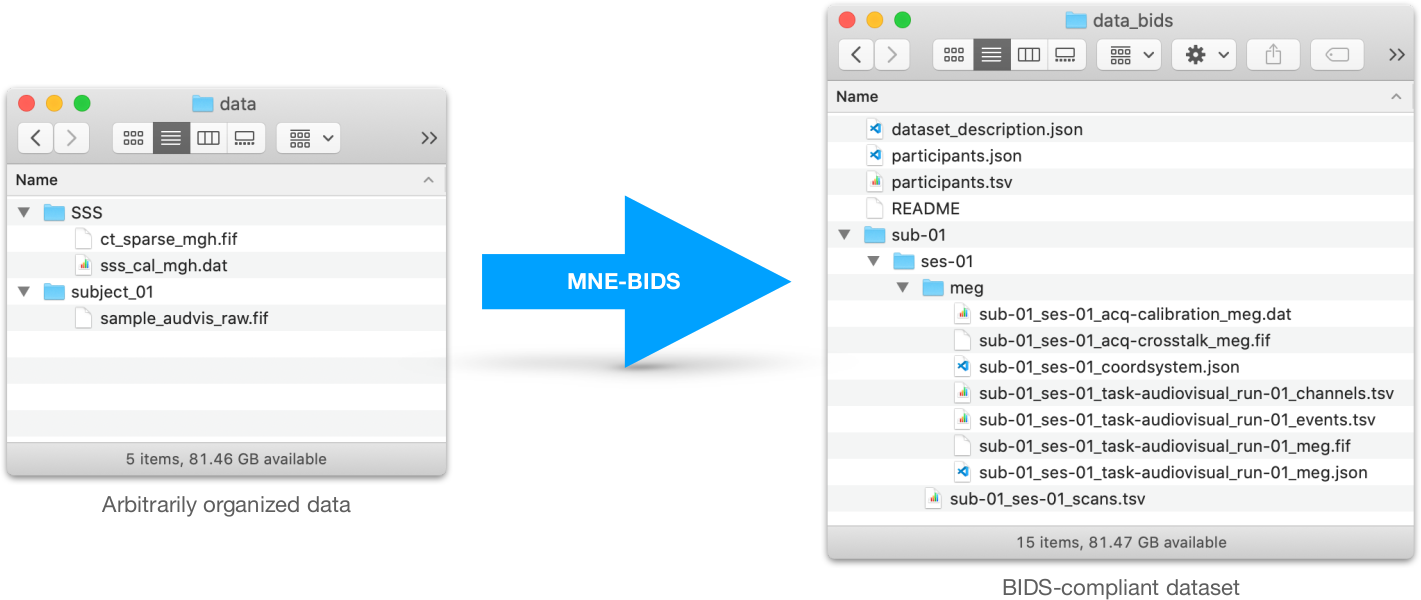
MNE-BIDS is a Python package that allows you to read and write BIDS-compatible datasets with the help of MNE-Python.
MNE-BIDS links BIDS and MNE-Python with the goal to make your analyses faster to code and more robust, and to facilitate data and code sharing with co-workers and collaborators.
Having your data in BIDS format also allows the use of automated pipelines, for example mne-bids-pipeline.
The documentation can be found under the following links:
- for the stable release
- for the latest (development) version
For any usage questions, please post to the MNE Forum.
Be sure to add the mne-bids tag to your question.
If you use MNE-BIDS in your work, please cite our publication in JOSS:
Appelhoff, S., Sanderson, M., Brooks, T., Vliet, M., Quentin, R., Holdgraf, C., Chaumon, M., Mikulan, E., Tavabi, K., Höchenberger, R., Welke, D., Brunner, C., Rockhill, A., Larson, E., Gramfort, A., & Jas, M. (2019): MNE-BIDS: Organizing electrophysiological data into the BIDS format and facilitating their analysis. Journal of Open Source Software, 4:1896. DOI: 10.21105/joss.01896
Please also cite one of the following papers to credit BIDS, depending on which data type you used: