-
Notifications
You must be signed in to change notification settings - Fork 32
Step 2: Align
Mark Harfouche edited this page Jun 20, 2018
·
16 revisions
In the Align step, TeraStitcher computes a series of displacements (one per layer: layers divide the volume along Z) for each pair of adjacent tiles.
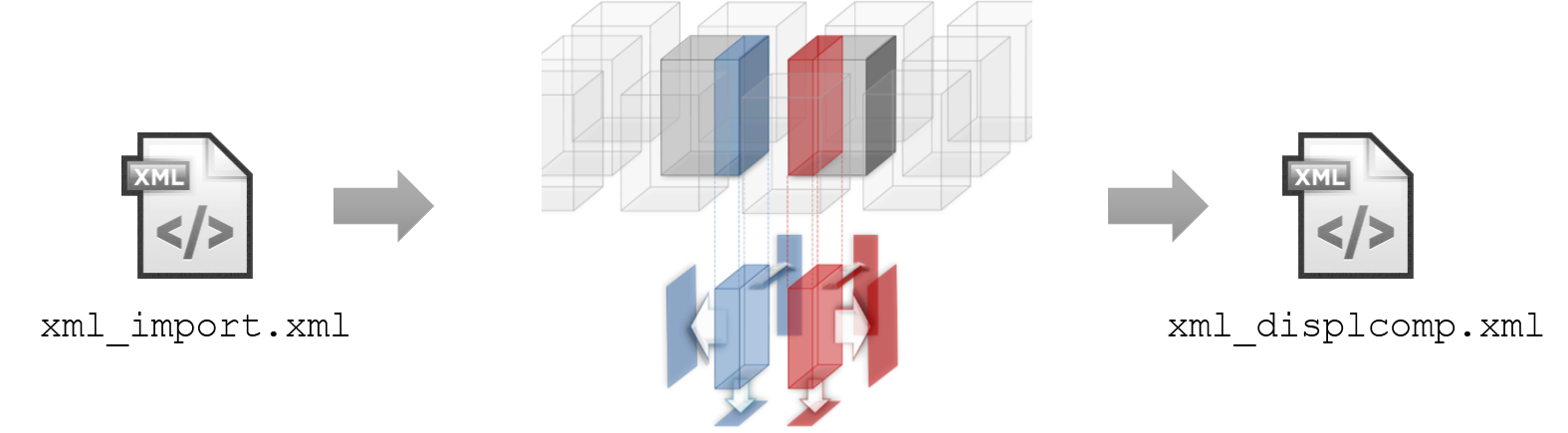
The algorithm used by default is called MIP-NCC and it is based on the Normalized Cross-Correlation applied on the three pairs of Maximum Intensity Projections extracted from the pair of tiles to be aligned.
- a volume that has been previously imported (see Import step) and the correspondent XML file descriptor produced.
- an installed TeraStitcher I/O plugin supporting your image file format. You can also write your own I/O plugin.
- absolute path of the XML descriptor (
--projin)
- output XML descriptor filepath (
--projout) - number of slices per layer (
--subvoldim) - data subset selection (
--R0--R1[--C0] (https://github.com/abria/TeraStitcher/wiki/User-Interface#--c0integer)--C1--D0--D1) - estimated tile overlap (
--oV--oH) - displacement search region (
--sV--sH--sD) - whether to compute images descriptors to enable subsequent SPIM artifacts removal (
--restoreSPIM--restoredir) - algorithm (
--algorithm) - image channel selection (
--imin_channel)
- XML descriptor saved at
--projoutcontaining a series of displacements for each pair of adjacent tiles.
terastitcher --displcompute --projin="C:/volume/xml_import.xml" --subvoldim=100
Wiki maintained by Alessandro Bria and Giulio Iannello.
- Quick Guide (pdf)
- TeraTools Guide (pdf)
- Stitching pipeline
- User Interface
- Supported volume formats
- Demo and batch scripts
- Built-in I/O plugins
- FAQ
