- Description
- Implementation and requirements
- Install
- Dependencies
- Documentation
- Citing ntEdit
- Credits
- How to run ntEdit
- Running ntEdit
- ntEdit polishing options
- Soft-mask option
- SNV mode
- VCF input option
- Test data
- Algorithm
- Output files
- License
ntEdit is a fast and scalable genomics application for polishing genome sequence assembly drafts. It simplifies polishing, variant detection and "haploidization" of gene and genome sequences with its re-usable Bloom filter design. Although it was originally designed as a general-purpose polishing tool, initally aimed at improving genome sequences by fixing base mismatches and frame shift errors with the help of more base-accurate short sequencing reads, ntEdit can also be used for polishing with long reads and to "finish" genome sequence assembly projects (refer to GoldPolish and the ntedit+sealer genome assembly finishing protocol, respectively).
We anticipate that ntEdit will find further applications in the rapid mapping of single nucleotide variants, as demonstrated below with the genome of SARS-CoV-2, the highly transmissible pathogenic coronavirus, and etiological agent of COVID-19. Additionally, for researchers delving into the intricate roots of genetic lineages within large cohort data, we encourage the utilization of ntRoot, an ancestry prediction framework built upon the ntEdit engine. It offers a comprehensive analysis of genetic heritage, employing sequence alignment-free algorithms to unveil ancestral connections and provide insights into genetic ancestry within diverse populations.
! NOTE: In v1.3.1 onwards, the parameter k is automatically detected from supplied Bloom filters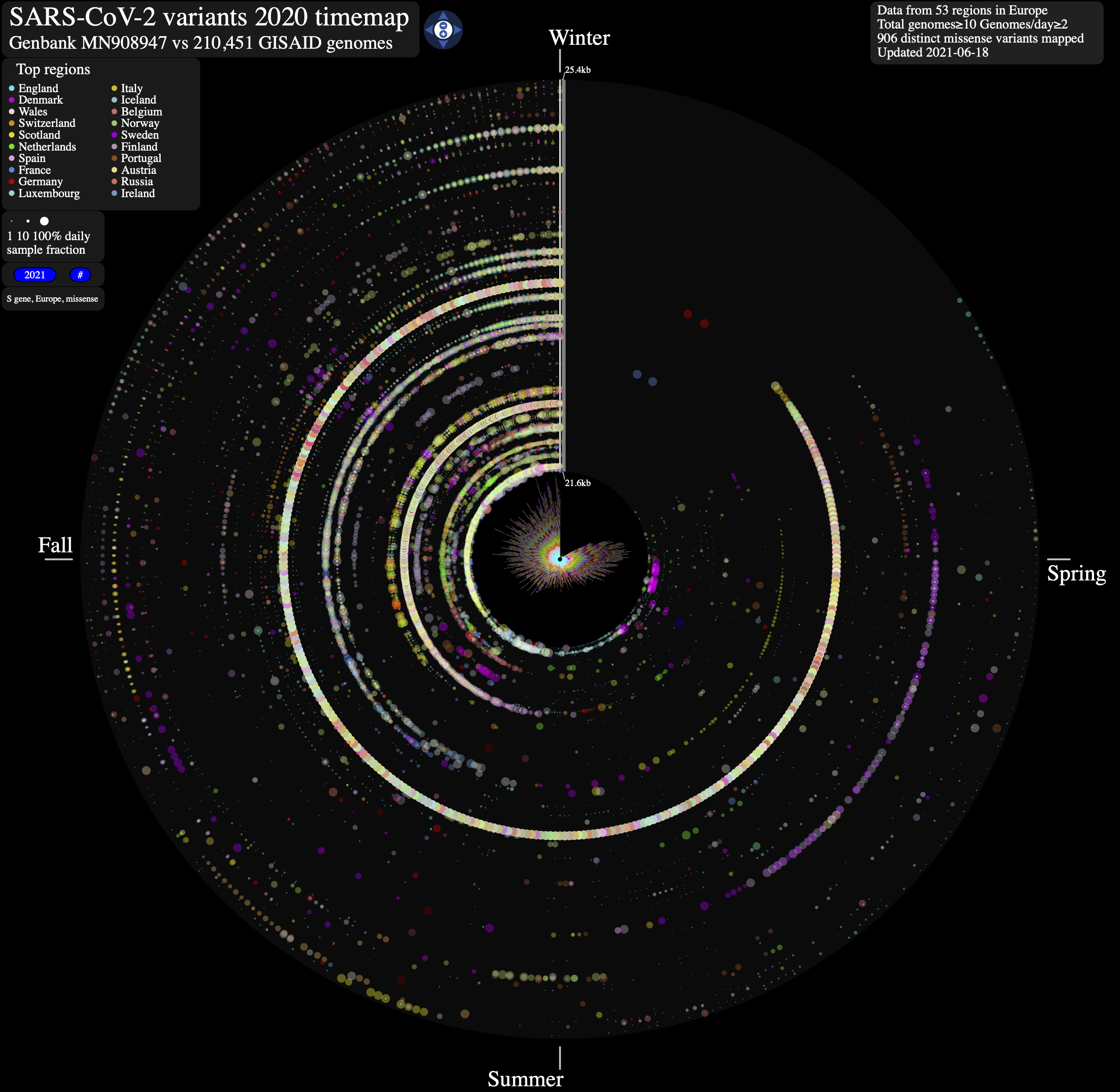
ntEdit v1.2.0 and subsequent versions are written in C++.
(We compiled with gcc 5.5.0)
Clone and enter the ntEdit directory.
git clone https://github.com/bcgsc/ntEdit.git cd ntEdit
Compile ntEdit.
meson setup build --prefix=/path/to/ntedit/install/dir cd build ninja install
- ntHits (v1.0.0+, https://github.com/bcgsc/nthits)
- BloomFilter utilities (provided in ./lib)
- kseq (provided in ./lib)
- meson
- ninja
- btllib
- snakemake
- python 3.9+
! NOTE: ntEdit v2.0.0+ IS ONLY compatible with ntHits release v1.0.0+We recommend installing ntEdit and its dependencies, using conda:
conda install -c bioconda ntedit
Refer to the README.md file on how to install and run ntEdit.
Our manuscript contains information about the software and its performance.
 This ISMB2019 poster contains additional information, benchmarks and results.
This ISMB2019 poster contains additional information, benchmarks and results.
Thank you for your 
If you use ntEdit in your research, please cite:
ntEdit: scalable genome sequence polishing
ntEdit: scalable genome sequence polishing. Warren RL, Coombe L, Mohamadi H, Zhang J, Jaquish B, Isabel N, Jones SJM, Bousquet J, Bohlmann J, Birol I. Bioinformatics. 2019 Nov 1;35(21):4430-4432. doi: 10.1093/bioinformatics/btz400.
The experimental data described in our paper can be downloaded from: http://www.bcgsc.ca/downloads/btl/ntedit/
ntedit (concept, algorithm design and prototype): Rene Warren
nthits / nthash: Hamid Mohamadi, Parham Kazemi
C++ implementation: Jessica Zhang, Rene Warren, Johnathan Wong
Integration tests: Murathan T Goktas
ntEdit workflow: Johnathan Wong and Lauren Coombe
General ntEdit usage:
run-ntedit --help
usage: run-ntedit [-h] {polish,snv} ...
ntEdit: Fast, lightweight, scalable genome sequence polishing and SNV detection & annotation
positional arguments:
{polish,snv} ntEdit can be run in polishing or SNV modes.
polish Run ntEdit polishing
snv Run ntEdit SNV mode (Experimental)
optional arguments:
-h, --help show this help message and exit
run-ntedit polish --help
usage: run-ntedit polish [-h] --draft DRAFT --reads READS [-i {0,1,2,3,4,5}] [-d {0,1,2,3,4,5,6,7,8,9,10}] [-x X] [--cap CAP] [-m {0,1,2}] [-a {0,1}] -k K
[-l L] [--cutoff CUTOFF] [--solid] [-t T] [-z Z] [-y Y] [-j J] [-X X] [-Y Y] [-v] [-V] [-n] [-f]
optional arguments:
-h, --help show this help message and exit
--draft DRAFT Draft genome assembly. Must be specified with exact FILE NAME. Ex: --draft myDraft.fa (FASTA, Multi-FASTA, and/or gzipped compatible),
REQUIRED
--reads READS Prefix of reads file(s). All files in the working directory with the specified prefix will be used for polishing (fastq, fasta, gz),
REQUIRED
-i {0,1,2,3,4,5} Maximum number of insertion bases to try, range 0-5, [default=5]
-d {0,1,2,3,4,5,6,7,8,9,10}
Maximum number of deletions bases to try, range 0-10, [default=5]
-x X k/x ratio for the number of k-mers that should be missing, [default=5.000]
--cap CAP Cap for the number of base insertions that can be made at one position[default=k*1.5]
-m {0,1,2} Mode of editing, range 0-2, [default=0] 0: best substitution, or first good indel 1: best substitution, or best indel 2: best edit
overall (suggestion that you reduce i and d for performance)
-a {0,1} Soft masks missing k-mer positions having no fix (1 = yes, default = 0, no)
-k K k-mer size, REQUIRED
-l L input VCF file with annotated variants (e.g., clinvar.vcf)
--cutoff CUTOFF The minimum coverage of k-mers in output Bloom filter [default=2, ignored if solid=True]
--solid Output the solid k-mers (non-erroneous k-mers), [default=False]
-t T Number of threads [default=4]
-z Z Minimum contig length [default=100]
-y Y k/y ratio for the number of edited k-mers that should be present, [default=9.000]
-j J controls size of k-mer subset. When checking subset of k-mers, check every jth k-mer [default=3]
-X X Ratio of number of k-mers in the k subset that should be missing in orderto attempt fix (higher=stringent) [default=0.5, if -Y is
specified]
-Y Y Ratio of number of k-mers in the k subset that should be present to accept an edit (higher=stringent) [default=0.5, if -X is specified]
-v Verbose mode, [default=False]
-V, --version show program's version number and exit
-n, --dry-run Print out the commands that will be executed
-f, --force Run all ntEdit steps, regardless of existing output files
run-ntedit snv --help
usage: run-ntedit snv [-h] [--reference REFERENCE] [--reads READS] [--genome GENOME [GENOME ...]] -k K [-l L] [--cutoff CUTOFF] [--solid] [-t T] [-z Z] [-y Y]
[-j J] [-X X] [-Y Y] [-v] [-V] [-n] [-f]
optional arguments:
-h, --help show this help message and exit
--reference REFERENCE
Reference genome assembly for SNV calling (FASTA, Multi-FASTA, and/or gzipped compatible), REQUIRED
--reads READS Prefix of input reads file(s) for variant calling. All files in the working directory with the specified prefix will be used for
polishing (fastq, fasta, gz)
--genome GENOME [GENOME ...]
Genome assembly file(s) for detecting SNV on --reference
-k K k-mer size, REQUIRED
-l L input VCF file with annotated variants (e.g., clinvar.vcf)
--cutoff CUTOFF The minimum coverage of k-mers in output Bloom filter [default=2, ignored if solid=True]
--solid Output the solid k-mers (non-erroneous k-mers), [default=False]
-t T Number of threads [default=4]
-z Z Minimum contig length [default=100]
-y Y k/y ratio for the number of edited k-mers that should be present, [default=9.000]
-j J controls size of k-mer subset. When checking subset of k-mers, check every jth k-mer [default=3]
-X X Ratio of number of k-mers in the k subset that should be missing in orderto attempt fix (higher=stringent) [default=0.5, if -Y is specified]
-Y Y Ratio of number of k-mers in the k subset that should be present to accept an edit (higher=stringent) [default=0.5, if -X is specified]
-v Verbose mode, [default=False]
-V, --version show program's version number and exit
-n, --dry-run Print out the commands that will be executed
-f, --force Run all ntEdit steps, regardless of existing output files
Example ntEdit command - polishing the draft ecoliWithMismatches001Indels0001.fa in solid mode using input reads my_reads_1.fq.gz and my_reads_2.fq.gz using a k-mer size of 55 and 48 threads
run-ntedit polish --draft ecoliWithMismatches001Indels0001.fa --reads my_reads -k 55 -t 48 --solid
run-ntedit polish --draft ecoliWithMismatches001Indels0001.fa --reads my_reads -k 55 -t 48 --cutoff 2
- For more advanced users, please see the help documentation for the
nteditexecutable, which has information about additional options- More information about the secondary Bloom filter mode is available on our wiki page
--solid modewill work well ONLY if you have sufficient read coverage (>30X). Otherwise, set the kmer coverage threshold to --cutoff 2 (>=20X) or --cutoff 1 (<20X)- solid mode will output non-error kmers, as determined by ntCard. Use this option only when you don't wish to set the threshold (--cutoff) manually
The ntEdit polishing option (or editing mode) is only used in polishing mode, and is controlled by -m
Mode 0: (default) ntEdit will try to substitute the last base of an incorrect k-mer with a different ATGC base. If that k-mer is found in the bloom filter and has enough subset support, ntEdit will then try the other substitution bases and then choose the best substitution fix. However, if the substituion was not found, then ntEdit will try all indels of max length (-i) and (-d) starting with that substitution base and make edit based on the first accepted indel. Mode 1: ntEdit will try to substitute the last base of an incorrect k-mer with a different ATGC base. If that k-mer is found in the bloom filter and has enough subset support, ntEdit will then try the other substitution bases and then choose the best substitution fix. However, if the substitution was nto found,t hen ntEdit will try all indels of max length (-i) and (-d) starting witht hat substitution base and make edit based on the best accepted indel. Mode 2: ntEdit will choose the best substitution or indel for each incorrect k-mer. Since this can be very computationally expensive because ntEdit tries every combination possible, it is recommended that you reduce (-i) and (-d).
*We recommend running ntEdit polishing in Mode 1 (or 0)
See https://github.com/bcgsc/ntedit_sealer_protocol and https://github.com/bcgsc/goldrush-edit for genome polishing pipelines that make use of this mode
Version 1.3.5 implements a new option (-a), which controls soft-masking (lower case) nucleotides in the supplied input [draft genome] sequence when its kmers are not found in the primary Bloom filter, and with no possible fix found in that filter (and optionally within a coverage slice provided by the secondary Bloom filter). This option is useful for flagging unpolished/unresolved genomic regions, those with no equivalent in the supplied Bloom filter(s). The nucleotide soft-masking effectively "paints a target" for other polishers/genome analysis software. This strategy is used in the ntedit_sealer_protocol and GoldPolish [a.k.a. goldrush-edit] (URLs above)
This mode can be useful for identifying unresolved genomic regions, those with no equivalent in the supplied Bloom filter(s). Version 1.3+ implements a new mode (`run-ntedit snv`) to help detect simple base variation in genome sequences. It works by overriding the kmer absence verification stage of ntEdit, effectively testing every base position for possible alternate k kmers. At the moment, ntEdit only reports possible base substitutions (no indels), along with the number of supported kmers (the latter is NOT a proxy for read/kmer coverage). In our tests on simulated (C. elegans, H. sapiens) and experimental (GIAB, HG001/HG004), we find k52/k55 (-j 3 -- see below) to give the best performance. Caveats: Variations occurring within 2*k are not reported. Because kmers are shorter and have less sequence context than reads and read pairs, kmer variations that occur within a genomic allele (intra allelic) may be reported. In order to minimize false discovery, we recommend using a secondary Bloom filter built with repeat kmers (see details on the -e Secondary Bloom filter option below). This option is provided as a convenience feature, implemented to do a quick and dirty variant detection analysis on large genomes. It is a basic presence/absence detector based on kmer subsets. For robust variant identification, we recommend statistically principled approaches. VCF output (v1.3.2+ _variants.vcf): We assume a diploid genome for reporting on the possible genotype (GT). Users working on polyploid genomes should chose to ignore the last two columns of the VCF file (ie. FORMAT INTEGRATION)
This handy option is used to supply a VCF input file to ntEdit, for cross-referencing base variants. For instance, users may wish to identify annotated clinical variants in their genomics datasets. For this, users would build Bloom filters with their read datasets using ntHits and run ntEdit in -s 1 mode, with the reference human genome as (-f) input. Note: it will also work in polishing mode on single nucleotide variants, but is of limited value since only homozygously divergent sites (i.e., with completely absent k-mers and k*k-mer) are reported in polishing mode.
We recommend the use of clinvar resources: https://www.ncbi.nlm.nih.gov/clinvar/ https://ftp.ncbi.nlm.nih.gov/pub/clinvar/vcf_GRCh38/ https://ftp.ncbi.nlm.nih.gov/pub/clinvar/vcf_GRCh38/clinvar_XXDATEXX.vcf.gz
Note: If you use clinvar, you MUST ensure you use GRCh38 AND that the chromosome IDs in the headers of your supplied (--reference) GRCH38 FASTA file matches that of clinvar's (e.g. >1 in FASTA and 1 in ClinVar VCF's #CHROM column). If you use any other VCF files as (-l) input, ensure consistency with FASTA headers.
example command:
run-ntedit snv --reference GRCh38.fa --reads HG004 -k 50 -t 48 -l clinvar_20230813.vcf
where the input reads have the prefix HG004
The SNV mode can also work on an input draft assembly:
run-ntedit snv --reference GRCh38.fa --genome HG004.asm.fa -k 50 -t 48 -l clinvar_20230813.vcf
The demo script will use the installed ntEdit binary. Please ensure that the ntEdit binary is in your PATH.
export PATH=/path/to/ntEdit:$PATH
Running the demo
Go to ./demo (cd demo)
run:
./runme.sh
ntEdit will polish an E. coli genome sequence with ~0.001 substitution error rate and ~0.0001 indel rate
Expected files will be:
ntedit_k25_changes.tsv
ntedit_k25_edited.fa
Compare with:
ecoli_ntedit_k25_changes.tsv
ecoli_ntedit_k25_edited.fa
 Sequence reads are first shredded into kmers using ntHits, keeping track of kmer multiplicity. The kmers that pass coverage thresholds (using ntHits, --cutoff option builds a filter with kmers having a coverage higher than cutoff) are used to construct a Bloom filter (BF). The draft assembly is supplied to ntEdit (--draft option, fasta file), along with the BF and sequences are read sequentially. Sequence strings are shredded into words of length k (kmers) at a specified value (-k option in versions before v1.3.1. In newer releases, k is detected automatically from the main Bloom filter) matching that used to build the BF, and each kmer from 5’ to 3’ queries the BF data structure for presence/absence (step 1). When a kmer is not found in the filter, a subset (Sk) of overlapping k kmers (defined by k over three, k/3) containing the 3’-end base is queried for absence (step 2). The subset Sk, representing a subsampling of k kmers obtained by sliding over 3 bases at a time over k bases, is chosen to minimize the number of checks against the Bloom filter. Of this subset, when the number of absent (-) kmers matches or exceeds a threshold defined by Sk- >= k/x (-x option), representing the majority of kmers in Sk, editing takes place (step 3 and beyond), otherwise step 1 resumes. In the former case, the 3’-end base is permuted to one of the three alternate bases (step 3), and the subset (Sk_alt) containing the change is assessed for Bloom filter presence (+). When that number matches or exceeds the threshold defined by Sk_alt+ >= k/y (-y option), which means the base substitution qualifies, it is tracked along with the number of supported kmers and the remaining alternate 3’-end base substitutions are also assessed (ie. resuming step 3 until all bases inspected). If the edit does not qualify, then a cycle of base insertion(s) and deletion(s) of up to –i and –d bases begins (step 4, -i option and step 5, -d option, respectively). As is the case for the substitutions, a subset of k kmers containing the indel change is tested for presence. If there are no qualifying changes, then the next alternate 3’-end base is inspected as per above; otherwise the change is applied to the sequence string and the next assembly kmer is inspected (step 1). The process is repeated until a qualifying change or until no suitable edits are found. In the latter case, we go back to step 1. When a change is made, the position on the new sequence is tracked, along with an alternate base with lesser or equal k kmer subset support, when applicable. Currently, ntEdit only tracks cases when edits are made (steps 3-5), and does not flag unedited, missing draft kmers (steps 1-2).
Sequence reads are first shredded into kmers using ntHits, keeping track of kmer multiplicity. The kmers that pass coverage thresholds (using ntHits, --cutoff option builds a filter with kmers having a coverage higher than cutoff) are used to construct a Bloom filter (BF). The draft assembly is supplied to ntEdit (--draft option, fasta file), along with the BF and sequences are read sequentially. Sequence strings are shredded into words of length k (kmers) at a specified value (-k option in versions before v1.3.1. In newer releases, k is detected automatically from the main Bloom filter) matching that used to build the BF, and each kmer from 5’ to 3’ queries the BF data structure for presence/absence (step 1). When a kmer is not found in the filter, a subset (Sk) of overlapping k kmers (defined by k over three, k/3) containing the 3’-end base is queried for absence (step 2). The subset Sk, representing a subsampling of k kmers obtained by sliding over 3 bases at a time over k bases, is chosen to minimize the number of checks against the Bloom filter. Of this subset, when the number of absent (-) kmers matches or exceeds a threshold defined by Sk- >= k/x (-x option), representing the majority of kmers in Sk, editing takes place (step 3 and beyond), otherwise step 1 resumes. In the former case, the 3’-end base is permuted to one of the three alternate bases (step 3), and the subset (Sk_alt) containing the change is assessed for Bloom filter presence (+). When that number matches or exceeds the threshold defined by Sk_alt+ >= k/y (-y option), which means the base substitution qualifies, it is tracked along with the number of supported kmers and the remaining alternate 3’-end base substitutions are also assessed (ie. resuming step 3 until all bases inspected). If the edit does not qualify, then a cycle of base insertion(s) and deletion(s) of up to –i and –d bases begins (step 4, -i option and step 5, -d option, respectively). As is the case for the substitutions, a subset of k kmers containing the indel change is tested for presence. If there are no qualifying changes, then the next alternate 3’-end base is inspected as per above; otherwise the change is applied to the sequence string and the next assembly kmer is inspected (step 1). The process is repeated until a qualifying change or until no suitable edits are found. In the latter case, we go back to step 1. When a change is made, the position on the new sequence is tracked, along with an alternate base with lesser or equal k kmer subset support, when applicable. Currently, ntEdit only tracks cases when edits are made (steps 3-5), and does not flag unedited, missing draft kmers (steps 1-2).
| Output files | Description |
|---|---|
| _changes.tsv | tab-separated file; ID bpPosition+1 OriginalBase NewBase Support k-mers (out of k/j) AlternateNewBase Alt.Support k-mers eg. U00096.3_MG1655_k12 117 A T 9 |
| _edited.fa | fasta file; contains the polished genome assembly |
| _variants.vcf | vcf file; contains variant calls |
note: ntEdit will polish input sequences in upper or lowercase bases. The case of bases in the input sequence IS preserved in the FASTA output, unless a fix is made by ntEdit (i.e., lower-case bases will remain lower-cased UNLESS a change is made).
ntEdit Copyright (c) 2018-current British Columbia Cancer Agency Branch. All rights reserved.
ntEdit is released under the GNU General Public License v3
This program is free software: you can redistribute it and/or modify it under the terms of the GNU General Public License as published by the Free Software Foundation, version 3.
This program is distributed in the hope that it will be useful, but WITHOUT ANY WARRANTY; without even the implied warranty of MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the GNU General Public License for more details.
You should have received a copy of the GNU General Public License along with this program. If not, see http://www.gnu.org/licenses/.
For commercial licensing options, please contact Patrick Rebstein prebstein@bccancer.bc.ca





