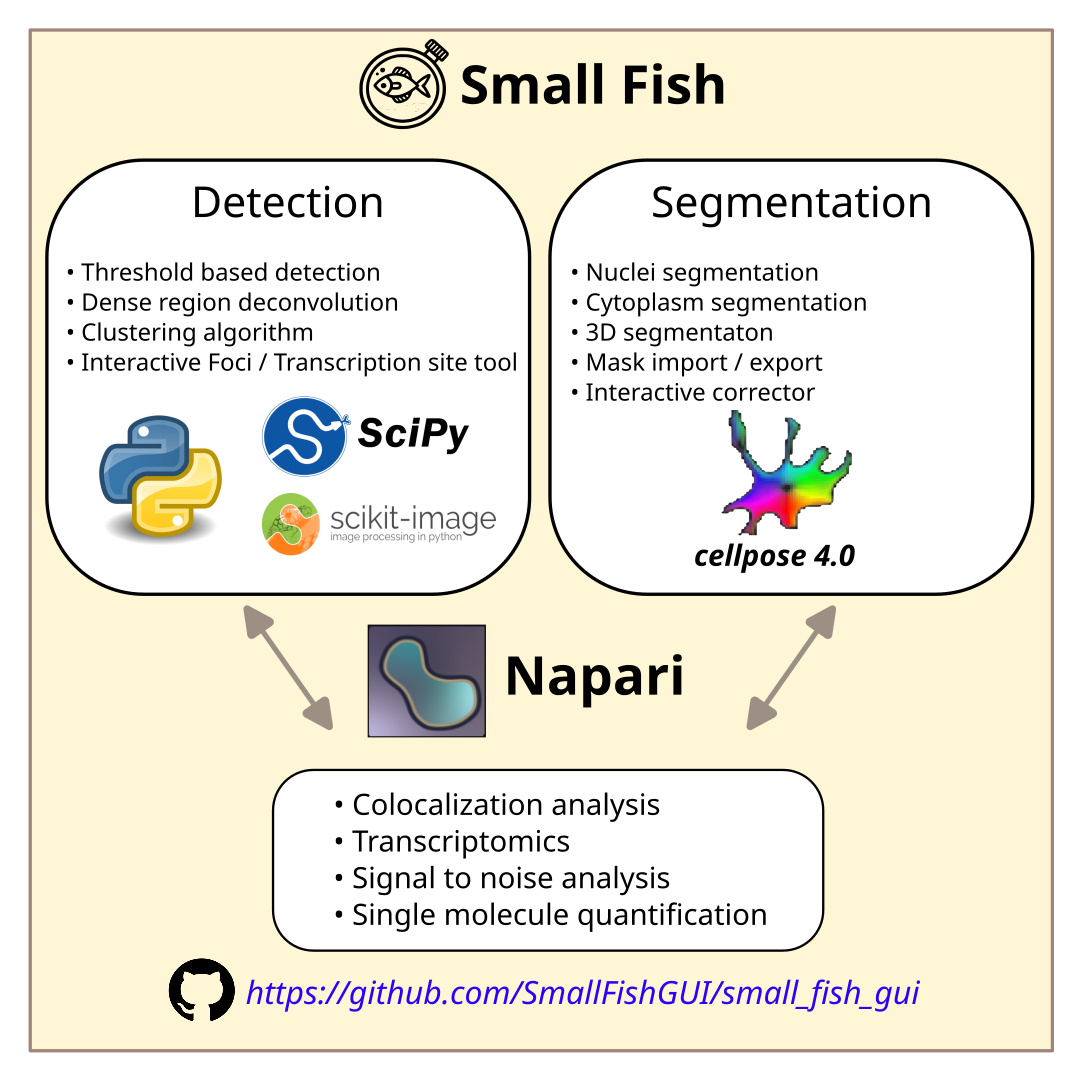
Small Fish is a python application for smFish image analysis. It provides a ready to use graphical interface to synthetize state-of-the-art scientific packages into an automated workflow. Small Fish is designed to simplify images quantification and analysis for people without coding skills.
Cell segmentation is peformed in 2D and 3D throught cellpose 4.0+(published work) : https://github.com/MouseLand/cellpose; compatible with your own cellpose models.
Spot detection is performed via big-fish a python implementation of FishQuant (published work) : https://github.com/fish-quant/big-fish.
The workflow is fully explained in the wiki ! Make sure to check it out.
✅ Single molecule quantification
✅ Transcriptomics
✅ Foci quantification
✅ Transcription sites quantification
✅ Nuclear signal quantification
✅ Signal to noise analysis
✅ Cell segmentation
✅ Multichannel colocalisation
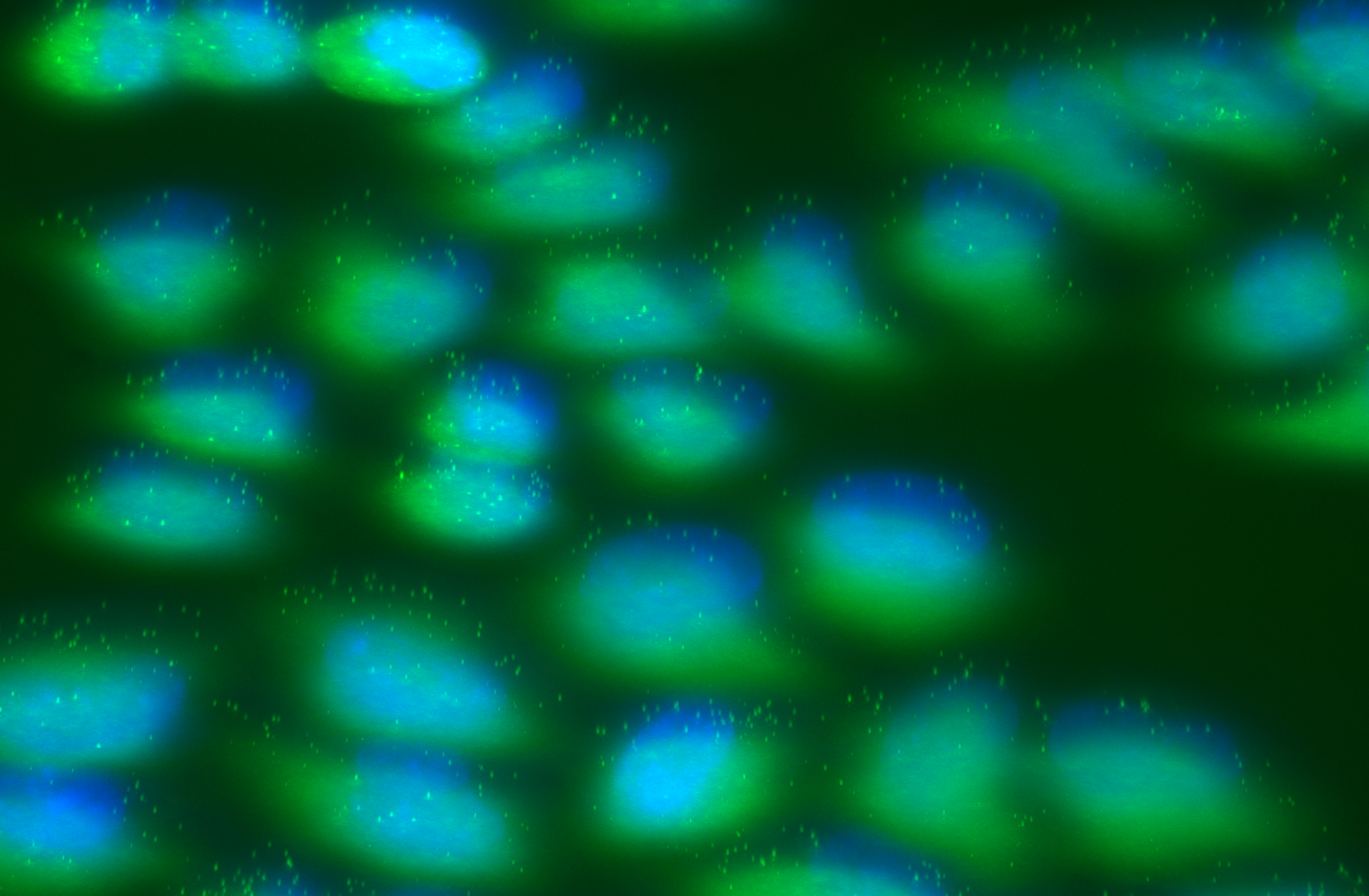
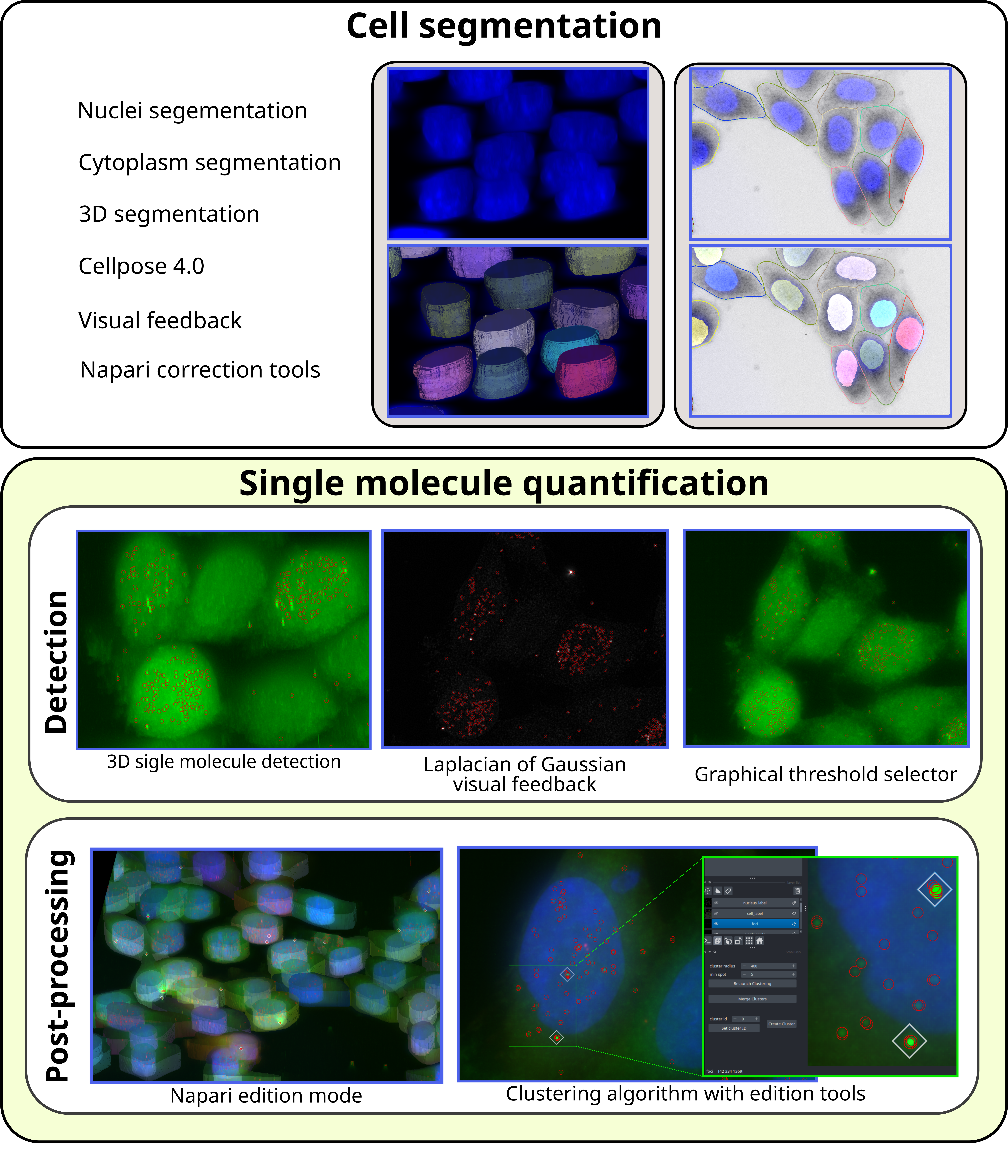
Raw 3D fish signal with dapi
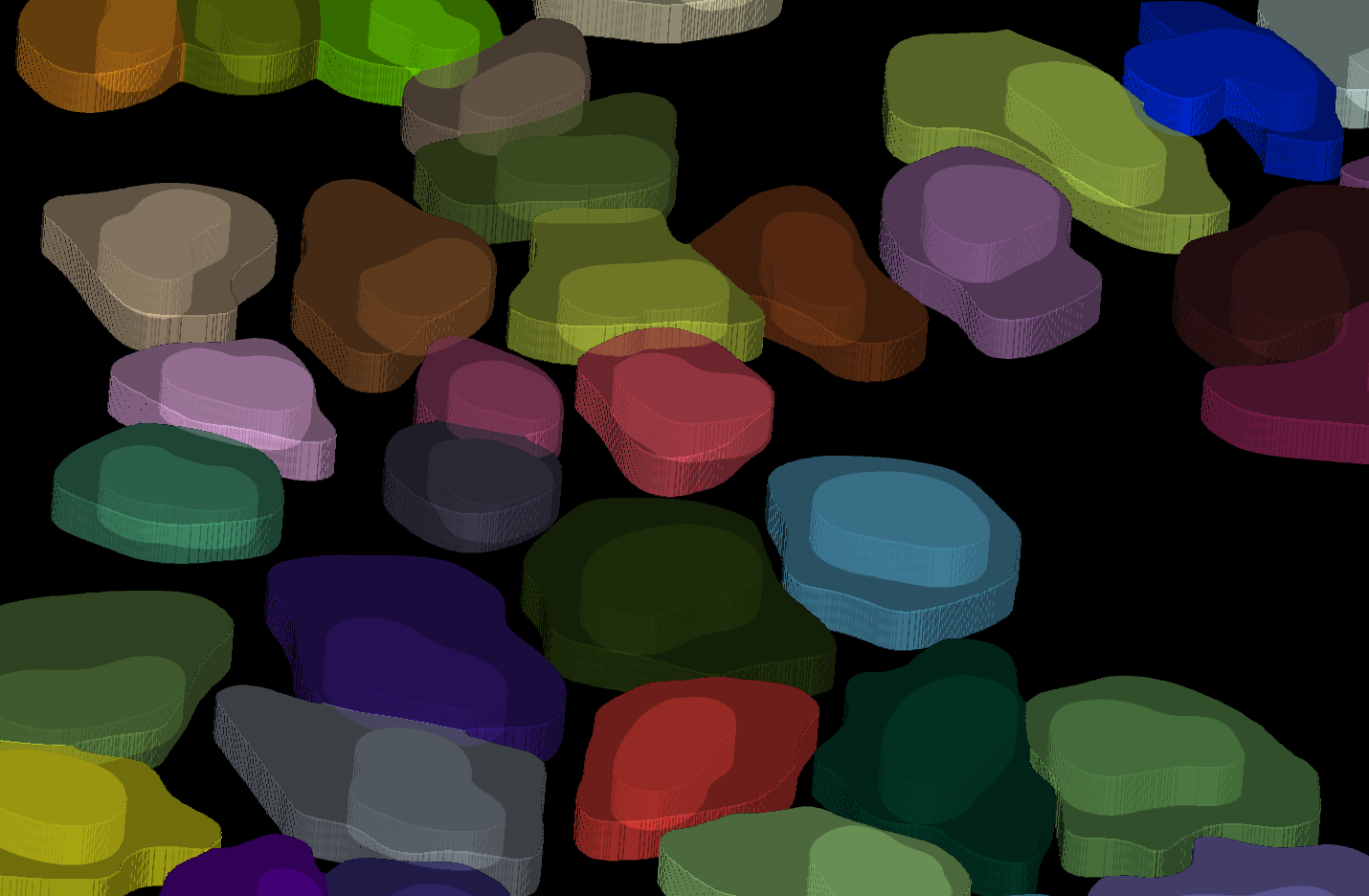
2D segmentation
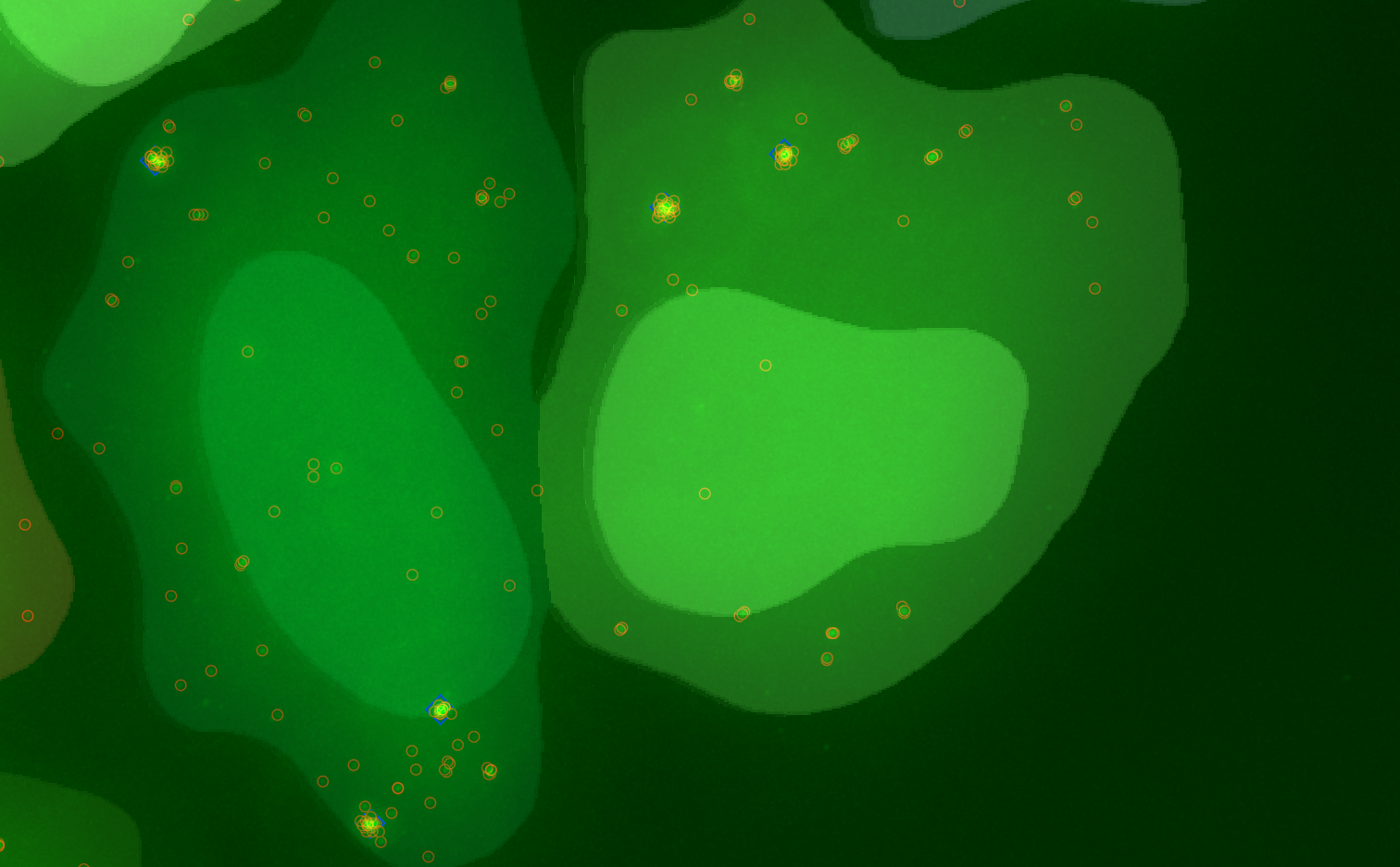
3D Spot detection
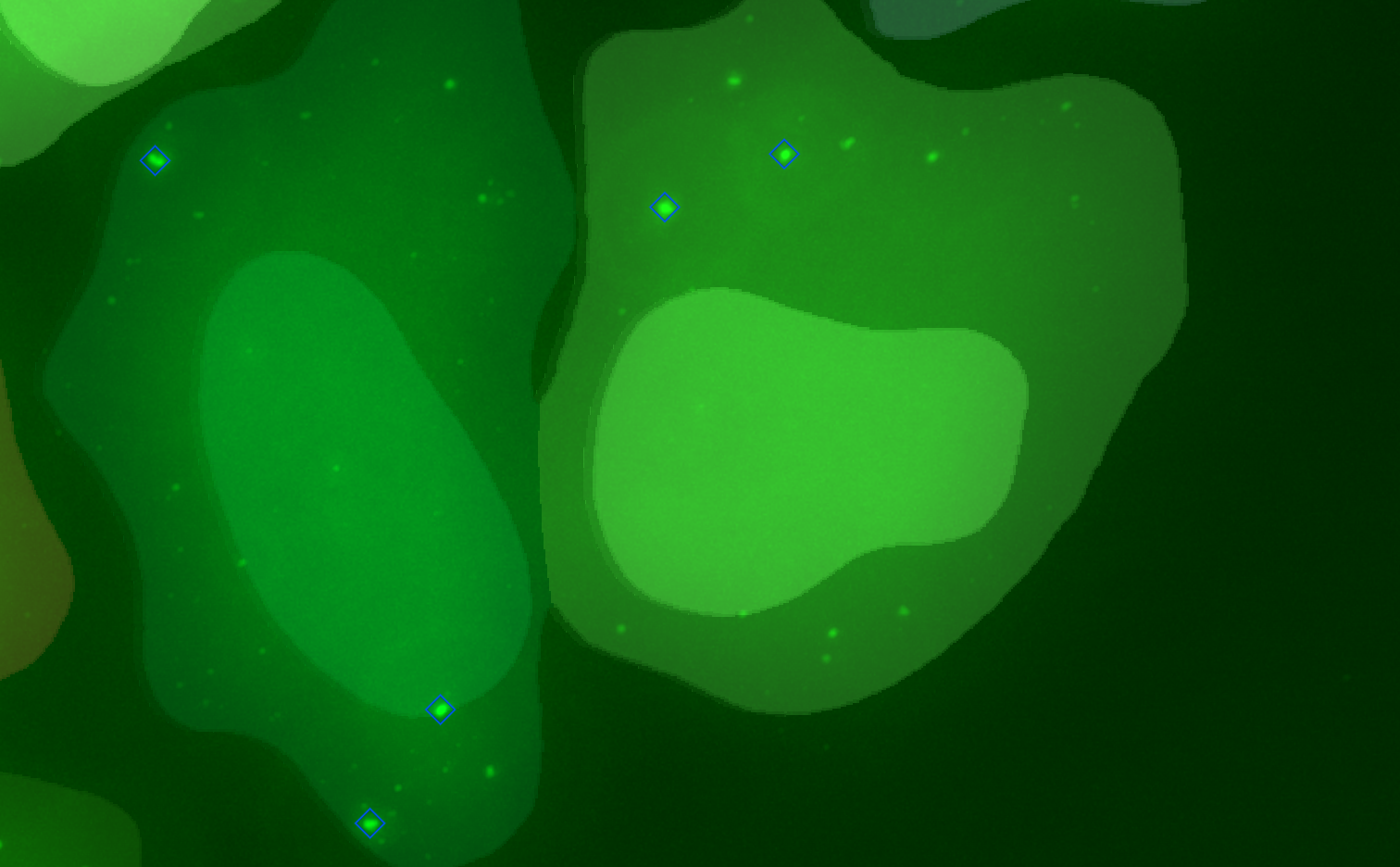
Cluster detection
Analysis can be performed either fully interactively throught a Napari interface or performed automatically through a batch processing allowing for reproducible quantifications.
If you don't have a python installation yet I would recommend the miniconda distribution; but any distribution should work.
It is higly recommanded to create a specific conda or virtual environnement to install small fish.
As of version 2.1.0 Small Fish runs on python 3.12.
If you are using conda or miniconda
conda create -n small_fish python=3.12
conda activate small_fishIf you are using venv, after installing the official python 3.12, python-venv and python-tk .
python3.12 -m venv python_env/small_fish
source python_env/small_fish/bin/activateThen download and install the small_fish package with :
pip install small_fish_guiA docker image is also available on Dockerhub.
If you can use the free version, I recommend using the docker desktop version and let it guide you. Otherwise you can install the command line version of Docker and run the following :
docker pull floric333/small_fish:latest
docker run -it --rm \
--env="DISPLAY=$DISPLAY" \
--env="QT_X11_NO_MITSHM=1" \
--volume="/tmp/.X11-unix:/tmp/.X11-unix:rw" \
floric333/small_fish:latestAs of Small Fish 2.0.1 it is highly recommanded to set up GPU with cellpose since its new model, CellposeSAM, is very heavy computationally even more when attempting 3D segmentation.
First of all, try to run small fish gpu without additional commands depending on your configuration it could work straight out of the box. If encoutering any issue try first the following :
Note: For MacOS users check first if your GPU is not already set up by launching a segmentation as the drivers might set up automatically upon installation.
pip install --index-url https://download.pytorch.org/whl/cu124 torch torchvision torchaudioIf running into additional problems please look at cellpose documentation.
First activate your python environnement :
conda activate small_fishThen launch Small fish :
python -m small_fish_guiYou are all set! Try it yourself or check the get started section in the wiki.