Online PHITS Tools documentation: lindt8.github.io/PHITS-Tools/
Purpose | Installation | Usage | CLI options | Automatic processing | Submodules | Testing/Issues/Contribution
This module is a collection of Python 3 functions that serve to automatically process, organize, and visualize output from the PHITS general purpose Monte Carlo particle transport code (and ease/expedite further analyses) and interfaces for utilizing these parsing/processing functions. PHITS can be obtained at https://phits.jaea.go.jp/.
Specifically, PHITS Tools seeks to be a universal PHITS output parser, supporting output from all tallies, both normal "standard" output as well as dump file outputs (in ASCII and binary formats), reading in the numeric data and metadata and storing them in Python objects for further use and analysis in Python. PHITS Tools is also coupled to the DCHAIN Tools module and can import it to process DCHAIN output when the main tally output parsing function is provided DCHAIN-related files. PHITS Tools also contains a number of functions for assisting in some types of further analyses. You can read more about how to use PHITS Tools and its output in its online documentation: lindt8.github.io/PHITS-Tools/
If you use/reference this code in a research paper, please cite this paper as follows:
Ratliff, H. N., (2025). The PHITS Tools Python package for parsing, organizing, and analyzing results from the PHITS radiation transport and DCHAIN activation codes. Journal of Open Source Software, 10(113), 8311, https://doi.org/10.21105/joss.08311
@article{Ratliff2025, doi = {10.21105/joss.08311}, url = {https://doi.org/10.21105/joss.08311}, year = {2025}, publisher = {The Open Journal}, volume = {10}, number = {113}, pages = {8311}, author = {Ratliff, Hunter N.}, title = {{The PHITS Tools Python package for parsing, organizing, and analyzing results from the PHITS radiation transport and DCHAIN activation codes}}, journal = {Journal of Open Source Software} } Install PHITS Tools:
pip install PHITS-Tools
Import PHITS Tools like any other Python module:
import PHITS_tools / from PHITS_tools import *
Note: To use the CLI/GUI, you must either use one of the commands PHITS-Tools/PHITS_tools/phits-tools/PHITS-Tools-GUI (more details further below) or execute the PHITS_tools.py module file with python. To find the installed location of the module file, execute: pip show PHITS-Tools -f
One may use the functions by first placing the PHITS_tools.py Python script into a folder in their PYTHONPATH system variable or in the active directory and then just importing them normally (import PHITS_tools / from PHITS_tools import *) or by executing the script python PHITS_tools.py with the PHITS output file to be parsed as the required argument (see python PHITS_tools.py --help for all CLI options) / without a file argument to be guided through with a GUI.
The short list of required package/library dependencies for PHITS Tools (and DCHAIN Tools) can be found in requirements.txt and installed by executing pip install -r requirements.txt.
To also be able to use the included submodules, download the latest source distribution (.zip/.tar.gz) from PyPI (includes only the minimum required source files) or the latest release (includes source plus example, tests, and docs), extract the contents, relocate the extracted directory of the package contents if desired, and make sure that directory is placed in your PYTHONPATH system variable (or active directory).
There are three main ways one can use this Python module:
- As an imported Python module
- In your own Python scripts, you can import this module (
import PHITS_tools/from PHITS_tools import *) and call its main functions or any of its other functions documented here.
- In your own Python scripts, you can import this module (
- As a command line interface (CLI)
- This module can be ran on the command line with the individual PHITS output file to be parsed (or a directory or PHITS input/phits.out file containing multiple files to be parsed) as the required argument. Execute
python PHITS_tools.py --helpto see all of the different options that can be used with this module to parse standard or dump PHITS output files (individually and directories containing them) via the CLI. This functionality can be used to automatically process all PHITS output whenever PHITS is ran; see the "Automatic processing at PHITS runtime" section further below.
- This module can be ran on the command line with the individual PHITS output file to be parsed (or a directory or PHITS input/phits.out file containing multiple files to be parsed) as the required argument. Execute
- As a graphical user interface (GUI)
- When the module is executed without any additional arguments,
python PHITS_tools.py, (or with the-gor--GUIflag in the CLI) a GUI will be launched to step you through selecting what "mode" you would like to run PHITS Tools in (STANDARD,DUMP,DIRECTORY, orINPUT_FILE), selecting a file to be parsed (or a directory containing multiple files to be parsed), and the various options for each mode.
- When the module is executed without any additional arguments,
Furthermore, if you have installed PHITS Tools via pip install PHITS-Tools, you will have access to the following
commands that can be used in place of python PHITS_tools.py for CLI usage in the terminal:
PHITS-Tools,PHITS_tools, andphits-tools
Additionally, an executable called PHITS-Tools-GUI is also created which can be ran (either in terminal or via
double-clicking it like any other executable file) to launch the GUI. If you prefer using the GUI, it may be convenient
to create a shortcut to this executable and place the shortcut somewhere easily accessible like your Desktop.
The executable will be located in your Python installation's "Scripts" folder; you can find it easily with where PHITS-Tools-GUI (Windows),
which PHITS-Tools-GUI (macOS/Linux), or python -c "import sysconfig; print(sysconfig.get_paths()['scripts'])" (all platforms).
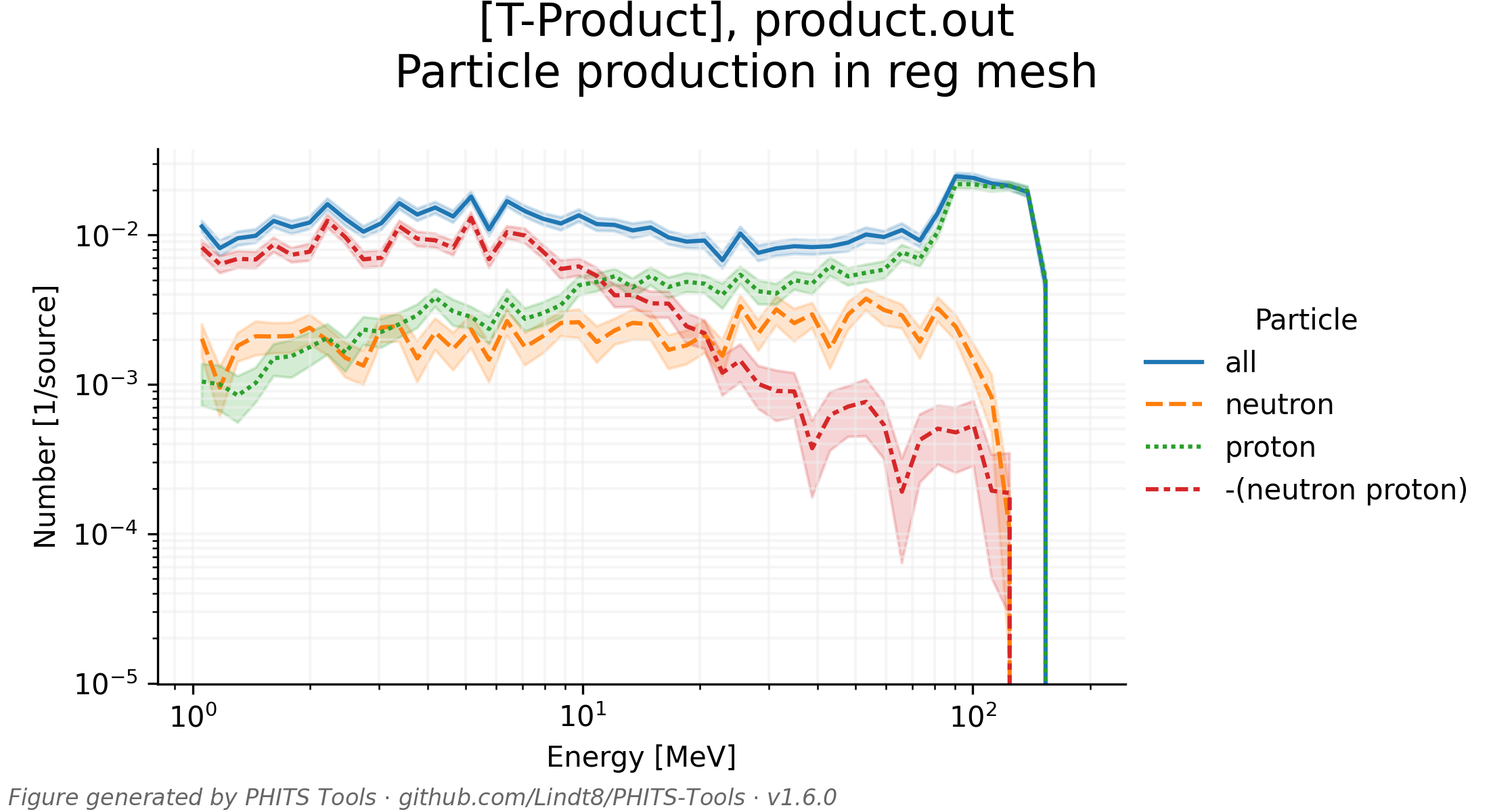
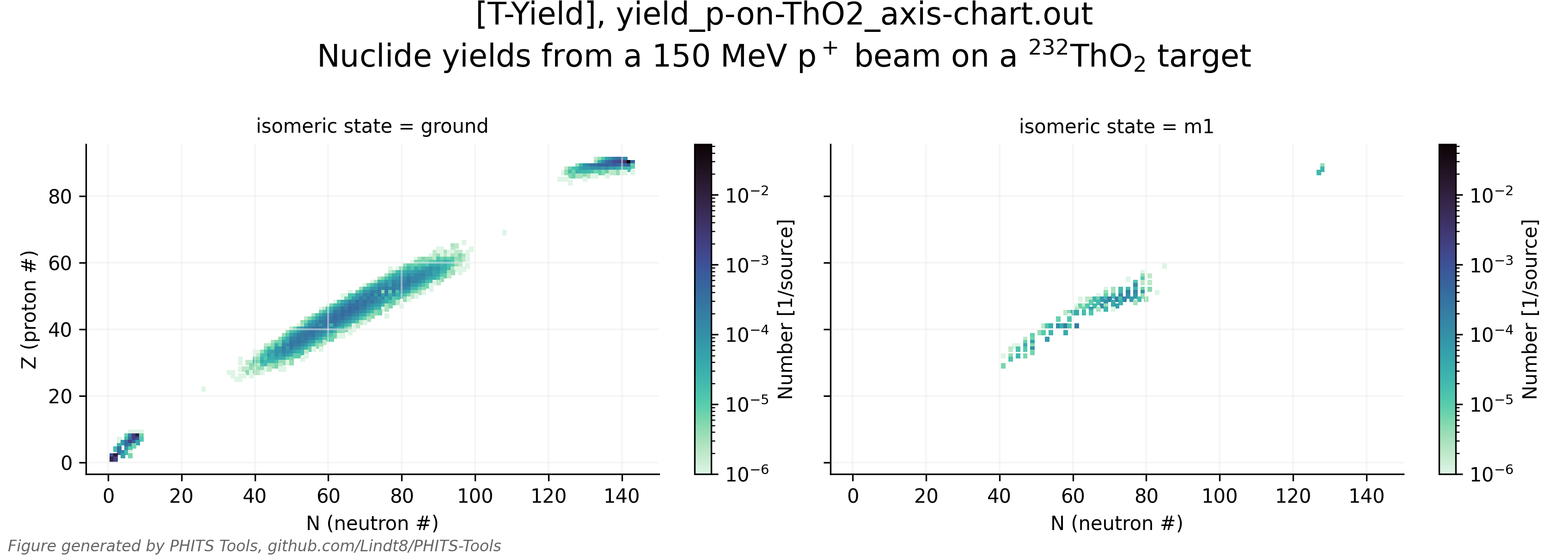
Aside from the main PHITS output parsing function parse_tally_output_file() for general tally output, the parse_tally_dump_file() function for parsing tally dump file outputs, and the parse_all_tally_output_in_dir() function for parsing all standard (and, optionally, dump) tally outputs in a directory or listed in a PHITS input/phits.out file, PHITS_tools.py also contains a number of other functions that may be of use for further analyses, such as tallying contents of dump files, rebinning historgrammed results, applying ICRP 116 effective dose conversion coefficients to scored particle fluences, and retrieving/modifying/adding PHITS-formatted [Material] section entries from a large database of materials (primarily from PNNL-15870 Rev. 1), among others. It also is capable of automatically creating plots of tally results, as showcased in test/test_tally_plots.pdf (view whole PDF here) and in the couple of example plots below.
 |
 |
|---|
The CLI and GUI options result in the parsed file's contents being saved to a pickle file, which can be reopened and used later in a Python script. (The pickle files produced when parsing "dump" output files are by default compressed via Python's built-in LZMA compression, indicated with an additional '.xz' file extension.) When using the main functions within a Python script which has imported the PHITS_tools module, you can optionally choose not to save the pickle files (if desired) and only have the tally output/dump parsing functions return the data objects they produce (dictionaries, NumPy arrays, Pandas DataFrames, and [only for dump outputs] lists of namedtuples / similarly functioning NumPy recarrays when saved to a pickle file) for your own further analyses.
Pictured below is the main PHITS Tools GUI window followed by the [DIRECTORY mode] GUI menu which shows all the options available not only for DIRECTORY mode but also for standard and dump tally output files, with the default options selected/populated.
The CLI principally serves to interface with the core three functions of PHITS Tools: parse_tally_output_file(), parse_tally_dump_file(), and parse_all_tally_output_in_dir(). The required file argument is checked to see if it is a directory or a file, and, if the latter, whether the -d option is used denoting a dump output file, otherwise defaulting to assuming it is a PHITS standard tally output file; then file and the relevant settings are sent to the corresponding main function. Explicitly, inclusion of the various CLI options have the following effects on the main functions' arguments and settings:
- Affecting all functions
fileis passed totally_output_filepath,path_to_dump_file, ortally_output_dirpath-skipsetsprefer_reading_existing_pickle = True(Falseif excluded)
parse_tally_output_file()(and passed to it viaparse_all_tally_output_in_dir())-npsetsmake_PandasDF = False(Trueif excluded)-nasetscalculate_absolute_errors = False(Trueif excluded)-lzmasetscompress_pickle_with_lzma = True(Falseif excluded)-posetsinclude_phitsout_in_metadata = True(Falseif excluded)-psetsautoplot_tally_output = True(Falseif excluded)
parse_tally_dump_file()(and passed to it viaparse_all_tally_output_in_dir())-dtells the CLI thatfileshould be processed as a dump file (if it's not a directory)-dvalspasses the provided sequence of values todump_data_sequence(Noneif excluded)-dbinspecifies that the file is binary (thatdump_data_number=len(dump_data_sequence)and is positive)-dnmaxpasses its value tomax_entries_read(Noneif excluded)-ddirsetsreturn_directional_info = True(Falseif excluded)-ddegsetsuse_degrees = True(Falseif excluded)-dnslsetssave_namedtuple_list = False(Trueif excluded)-dnspsetssave_Pandas_dataframe = False(Trueif excluded)-dmaxGBpasses its value tosplit_binary_dumps_over_X_GB(20GB if excluded)-dsplitpasses its value tomerge_split_dump_handling(0if excluded)
parse_all_tally_output_in_dir()exclusively-rsetsinclude_subdirectories = True(Falseif excluded)-fprepasses its value tooutput_file_prefix(''if excluded)-fsufpasses its value tooutput_file_suffix('.out'if excluded)-fstrpasses its value tooutput_file_required_string(''if excluded)-dsetsinclude_dump_files = True(Falseif excluded)-dnmmpisetsdump_merge_MPI_subdumps = False(Trueif excluded)-dndmpisetsdump_delete_MPI_subdumps_post_merge = False(Trueif excluded)-msetsmerge_tally_outputs = True(Falseif excluded)-smosetsmerge_tally_outputs = True(Falseif excluded),save_output_pickle = False(Trueif excluded), andsave_pickle_of_merged_tally_outputs = True(Noneif excluded)-pasetsautoplot_all_tally_output_in_dir = True(Falseif excluded)
Below is a picture of all of these options available for use within the CLI.
PHITS Tools can be used to automatically process the output of every PHITS run executed with the "phits.bat" and "phits.sh" scripts found in the "phits/bin/" directory of your PHITS installation. To do this, first you must identify the location of your "PHITS_tools.py" file. If using the file directly downloaded from GitHub, this should be in a location of your choosing. If you installed PHITS Tools via pip install PHITS-Tools, you can find its location with pip show PHITS-Tools -f. Once you have identified the location of PHITS_tools.py, for example "/path/locating/PHITS_Tools/PHITS_tools.py", you can add the following line to your PHITS execution script:
On Windows, using "phits/bin/phits.bat":
- Scroll down toward the bottom of the script, to the section with the line
rem - Your file processing starts here. - After the if statement (right before the
rem - Your file processing ends hereline), insert a new line with the following command: python "C:\path\locating\PHITS_Tools\PHITS_tools.py" "%%~nxF" -po -m -d -ddir -ddeg -lzma -p -pa- Or, if PHITS Tools was installed with
pip,python "C:\path\locating\PHITS_Tools\PHITS_tools.py"can be replaced withphits-toolsas:phits-tools "%%~nxF" -po -m -d -ddir -ddeg -lzma -p -pa
On Linux/Mac, using "phits/bin/phits.sh":
- Scroll down toward the bottom of the script, to the section titled
# Run PHITS - On the line after the end of the if statement
fi, add the following command: python "/path/locating/PHITS_Tools/PHITS_tools.py" $1 -po -m -d -ddir -ddeg -lzma -p -pa- Or, if PHITS Tools was installed with
pip,python "/path/locating/PHITS_Tools/PHITS_tools.py"can be replaced withphits-toolsas:phits-tools $1 -po -m -d -ddir -ddeg -lzma -p -pa
(Of course, if necessary, replace "python" with however you typically call python in your environment, e.g. py, python3, etc.)
Adding this line causes the following to happen:
- After PHITS finishes running normally, the PHITS input file is passed to PHITS Tools.
- Since it is a PHITS input file, the CLI will have
parse_all_tally_output_in_dir()handle it, in [INPUT_FILE mode] - The input file and its produced "phits.out"-type file are scanned for output files from active tallies, including those inputted via the PHITS insert file function
infl:{}too (usingextract_tally_outputs_from_phits_input()andparse_phitsout_file()).- This will include any dump files (
-d) if present. - When the "phits.out" file (
file(6)in the PHITS input [Parameters]) is parsed, its metadata—including the PHITS input echo—will be saved to a .pickle file, compressed with LZMA (-lzma) and with the extra ".xz" extension.
- This will include any dump files (
- Then, the standard tally outputs are processed. For each standard tally output:
- The
parse_tally_output_file()function processes the tally output, producing atally_outputdictionary with keys for the produced NumPy array, Pandas DataFrame, and metadata dictionary. - The
tally_output['tally_metadata']dictionary will have the phits.out metadata dictionary added to it (-po) under the'phitsout'key. - This
tally_outputdictionary object is saved to a .pickle file, compressed with LZMA (-lzma) and with the extra ".xz" extension. - A plot, saved in PDF and PNG formats, of the tally's output is generated (
-p) byautoplot_tally_results(). - These files will share the same name as the tally output file, just with different extensions.
- The
- Then, any encountered tally dump files are processed (
-d). For each tally dump output file:- The
parse_tally_dump_file()function processes the dump file, automatically extracting the dump parameters from its parent tally's output file. - Provided direction vector information (u,v,w) is present, extra directional information is calculated (
-ddir), using degrees for the unit of calculated angles (-ddeg) - The produced named tuple list and Pandas DataFrame are saved as two separate LZMA-compressed pickle files.
- The
- Then, a merged (
-m) dictionary object containing all of thetally_outputdictionaries for each standard tally output processed is produced.- The dictionary is keyed with the
fileparameter of each tally in the PHITS input, with the values being the correspondingtally_outputdictionaries. - This merged dictionary object is saved to a pickle file sharing the same name as the PHITS input file but ending in "_ALL_TALLY_OUTPUTS.pickle", compressed with LZMA (
-lzma) and with the extra ".xz" extension.
- The dictionary is keyed with the
- Then, a PDF containing plots from all standard tally outputs (
-pa) is generated withautoplot_tally_results().- This PDF of plots is saved to a file sharing the same name as the PHITS input file but ending in "_ALL_TALLY_OUTPUTS_PLOTTED.pdf"
You can edit the flags provided to the CLI for your desired default behavior. For instance, to only save the pickle file of the merged output (rather than for every tally output too), replace -m with -smo. And to not bother with creating a merged output and only save the outputs for each individual tally, just omit -m. Given that the plotting is easily the slowest part of the execution of PHITS Tools in most cases, it may be desirable to omit the -p and/or -pa flags to not save plots of the tally outputs individually or all together in a single PDF, respectively. Since dump files can be very large and sometimes solely created for reusage by PHITS (e.g., with a s-type=17 [Source] section), it may also be desirable to exclude dump files from automatic processing by omitting -d. As an example, a more "minimal" automatic processing would result from:
- Windows:
python "C:\path\locating\PHITS_Tools\PHITS_tools.py" "%%~nxF" -po -smo -lzma -pa - Linux/Mac:
python "/path/locating/PHITS_Tools/PHITS_tools.py" $1 -po -smo -lzma -pa
This would only create the ".pickle.xz" file of the merged standard tally outputs and the PDF containing all of their plots together, skipping any processing of dump files.
Similarly, PHITS Tools can be used to automatically process output from the DCHAIN code, utilizing an import of the DCHAIN Tools module.
On Windows, using "phits/dchain-sp/bin/dchain.bat":
- Scroll down toward the bottom of the script, to the section with the line
rem - Your file processing ends here. - Right above that line (before the
goto :continue), insert a new line with the following command: python "C:\path\locating\PHITS_Tools\PHITS_tools.py" "%%~nxF" -po -lzma- (or, if installed with
pip)phits-tools "%%~nxF" -po -lzma
- (or, if installed with
On Linux/Mac, using "phits/dchain-sp/bin/dchain.sh":
- Scroll down toward the bottom of the script, right before the line with
echo ' end of dchain ' - On the line after the end of the if statement
fi, add the following command: python "/path/locating/PHITS_Tools/PHITS_tools.py" ${jnam} -po -lzma- (or, if installed with
pip)phits-tools ${jnam} -po -lzma
- (or, if installed with
This will create a ".pickle.xz" file (with the same basename as the DCHAIN input file) of the processed DCHAIN outputs, as a dictionary object, with contents as described in the documentation for parse_tally_output_file() under the "[T-Dchain] special case" section, also including an entry with a dictionary of information on the corresponding PHITS run via its "phits.out" file (if found).
The PHITS Tools package consists of one main module (PHITS_tools.py) and two submodules,
listed below with their separate documentation linked. If you have installed the package via pip install PHITS-Tools,
these should work and be accessed as shown without any additional configuration required.
- DCHAIN Tools submodule:
dchain_tools.pyPHITS_tools.dchain_toolssubmodule documentation- Can be accessed with:
from PHITS_tools import dchain_tools - This submodule provides more customized processing of DCHAIN outputs.
- Monte Carlo materials management submodule:
manage_mc_materials.pyPHITS_tools.manage_mc_materialssubmodule documentation- Can be accessed with:
from PHITS_tools import manage_mc_materials - This submodule provides tools for building/customizing your own databases of materials used in Monte Carlo simulations.
I have extensively tested this module with a rather large number of PHITS output files with all sorts of different geometry settings, combinations of meshes, output options, and other settings to try to capture as a wide array of output files as I could (including the ~300 output files within the phits/sample/ and phits/recommendation/ directories included in the distributed PHITS release, which can be tested in an automated way with test/test_PHITS_tools.py in this repository, along with a large number of supplemental variations to really test every option I could think of), but there still may be some usage/combinations of different settings I had not considered that may cause PHITS Tools to crash when attempting to parse a particular output file. If you come across such an edge case—a standard PHITS tally output file that causes PHITS Tools to crash when attempting to parse it—please submit it as an issue and include the output file in question and I'll do my best to update the code to work with it! Over time, hopefully all the possible edge cases can get stamped out this way. :)
Likewise, if you have any questions or ideas for improvements / feature suggestions, feel free to submit them as an issue. If you would like to contribute a new function or changes to any existing functions, feel free to fork this repository, make a new branch with your additions/changes, and make a pull request. (GitHub has a nice short guide on this process.)
For more information on contributing and running tests yourself, please check out CONTRIBUTING.md, tests/ for unit tests, and test/ for integration/functional tests.
If using [T-Dchain] in PHITS and/or the DCHAIN-PHITS code, the DCHAIN Tools repository contains a separate Python module for parsing and processing that related code output. While PHITS Tools will import and use DCHAIN Tools if provided with DCHAIN-related files, direct usage of DCHAIN Tools may be desired if you want greater control of the various output parsing options within it or want to make use of some of its useful standalone functions. All of these functions are documented online at lindt8.github.io/DCHAIN-Tools/.
DCHAIN Tools is distributed as a submodule of PHITS Tools. If installing PHITS Tools via pip install PHITS-Tools, you can access DCHAIN Tools with from PHITS_tools import dchain_tools / from PHITS_tools.dchain_tools import *. If installing PHITS Tools manually, see dchain_tools.py in the DCHAIN-Tools directory/submodule link.
These functions are tools I have developed over time to speed up my usage of PHITS; they are not officially supported by the PHITS development team. All of the professionally-relevant Python modules I have developed are summarized here, and more general information about me and the work I do / have done can be found on my personal webpage.







