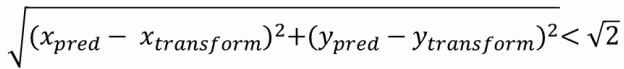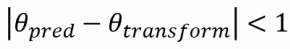In order to propose an accurate diagnosis during the longitudinal monitoring of the same region in magnetic resonance imaging, it is important that the same acquisition volumes over time are the same to allow a follow-up study.
To carry out an MRI acquisition of the brain, the operator must first define the region of interest to be visualized as well as the orientation of the cutting planes. To do this, it first performs low-resolution acquisitions through fast MRI sessions. This step is done manually. However, because of inter- and intra-operator variability, differences in positioning and orientation occur from one week to the next, making the diagnosis delicate or false.
Based on the low-resolution localizer images, the operator schedules a complete and detailed scan of the brain. This complete acquisition takes much longer. Therefore, it is essential that the positioning made with the location images is correct in order to obtain comparable acquisitions for longitudinal monitoring, and not having to relaunch an exam again.
The goal of the project is to establish a proof of feasibility of an automatic determination of the acquisition plans in cerebral MRI of the small animal under Python
By the time this project was done, there was no method to determine the acquisition plans for small animals, only for humans. The reason is that small animals have less water in their brains, making the acquisition image less informative and more noisy.
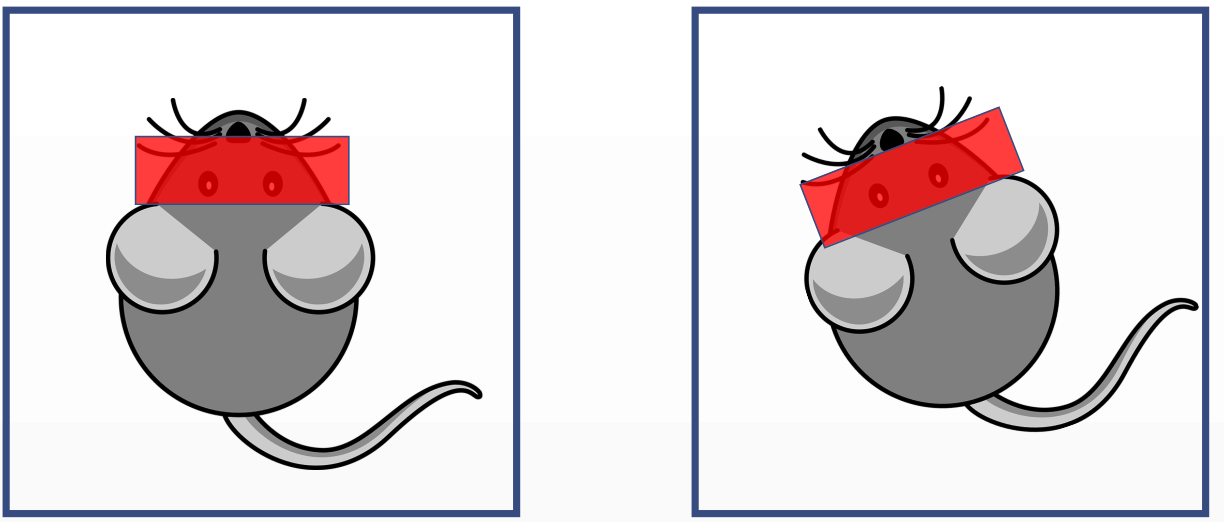
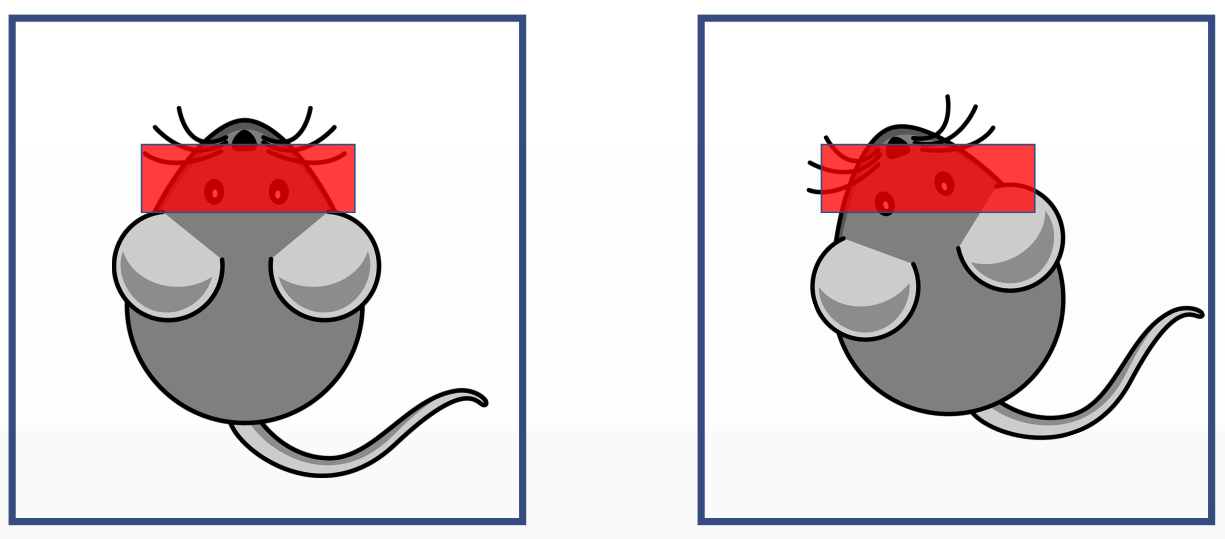
The small animal is placed in the MRI machine at week 0 and he is put back on the machine at week X to do a follow-up exam.
The acquisition volume would be wrong. To have have a compatible volume, the rotation and translation of the small animal inside the machine with relation to the former exam has to be fed to the machine so it can correct itself.
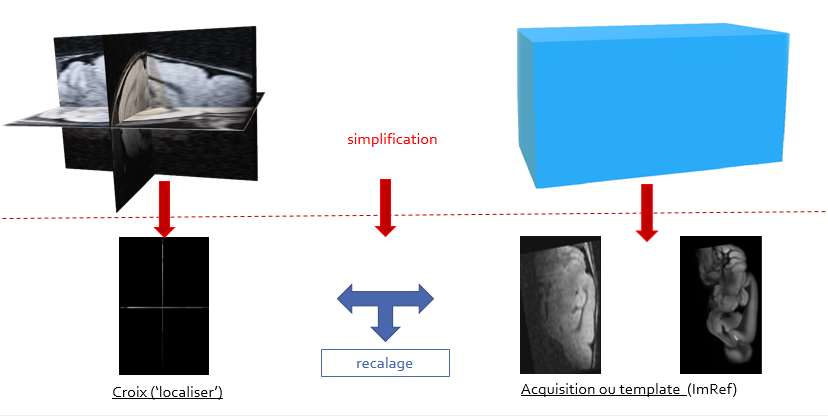
To obtain an MRI image at week x of the same volume as at week x-1, we first launch the rapid acquisition of a low resolution image composed of each of the three plans of the volume. This image will be called the ‘localizer’. We will readjust this last with the 3D volume (in blue on the right on the diagram) previously acquired, which will give the translation and rotation factors (according to x, y and theta) to enter into the software MRI positioning.
There are two ways the dataset is created. One is generate it using the previously volume acquired (intra-patient method).This method is not ideal as it requires a dataset generation and training of a model for each studied animal. The other is using an atlas image. It is an image created from the average of many other images of a brain. The dataset and model using the atlas is created only once, and all registrations will comply to the positioning relative to the atlas.
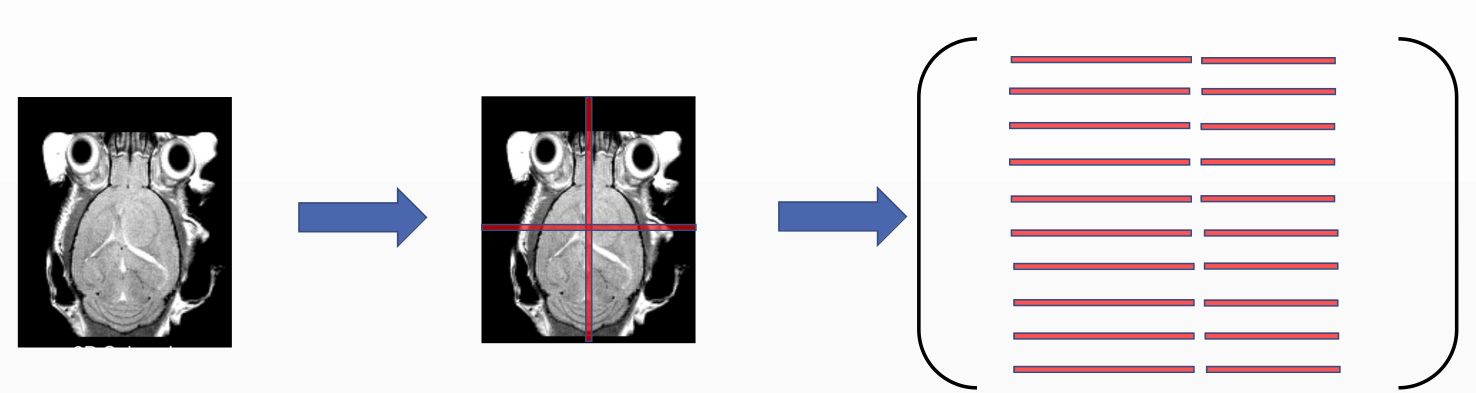
In our simplified 2D problem, the dataset creation is as follows: one image from the volume is taken and transformed along the x an y axis and rotated from the center. The chosen transformation was -30 to 30 pixels on x and y axis with a step of 1 pixel and -30 to 30 degrees with 1 theta step. All combination gives a dataset issued from 60x60x60 = 216000 images. From each image, the intensity values of the mid row and mid column are extracted and stored in the dataset matrix that later on is fed to the regression model.
To access the accuracy of the algorithm, we run a 1000 tests. At each test, the image is transformed with random parameters and the model tries to predict the transformation. It is considered a success if:
and
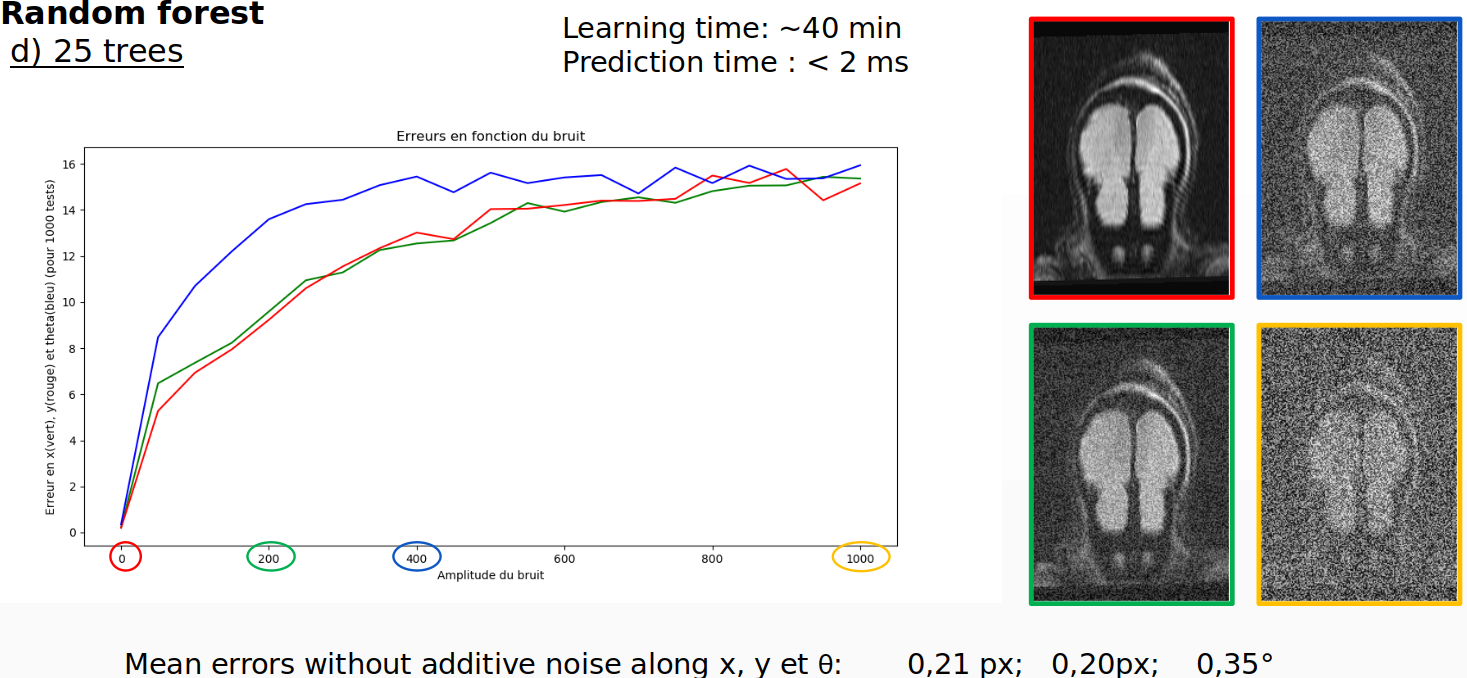
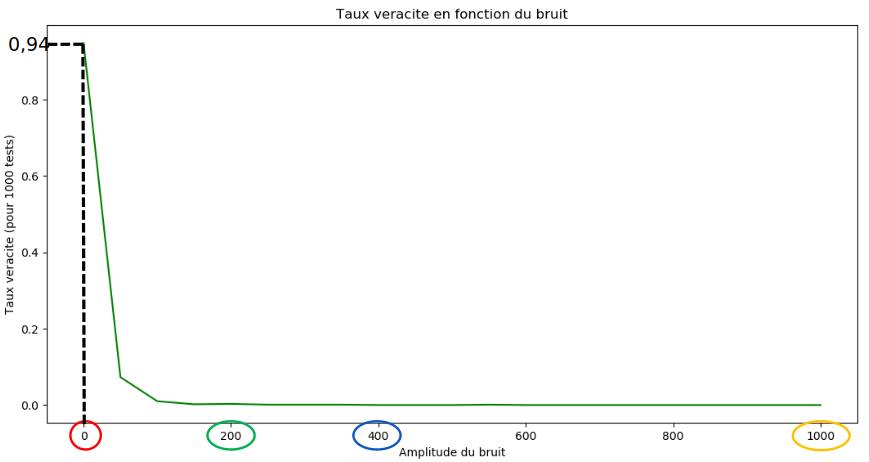
After the accuracy is accessed for this image, the accuracy is then accessed when noise is progressively added to the image. The values on the image range from -100 to 1200.
For the random forest regression with 25 trees trained on the acquisition, the success rate is 94%, the learning time is 40 minutes and prediction time is 2 milliseconds.
However, when the model is trained on the atlas image, the accuracy is virtually 0%. That means random forest could be considered for intra-patient exams. What make it impractical is the large learning time of 40 minutes and the model that weights 1GB. If more than just one image of the training volume would be used to train the model (3D problem) and considering now the two extra rotation along the other axis, the memory required for the model would easily explode.
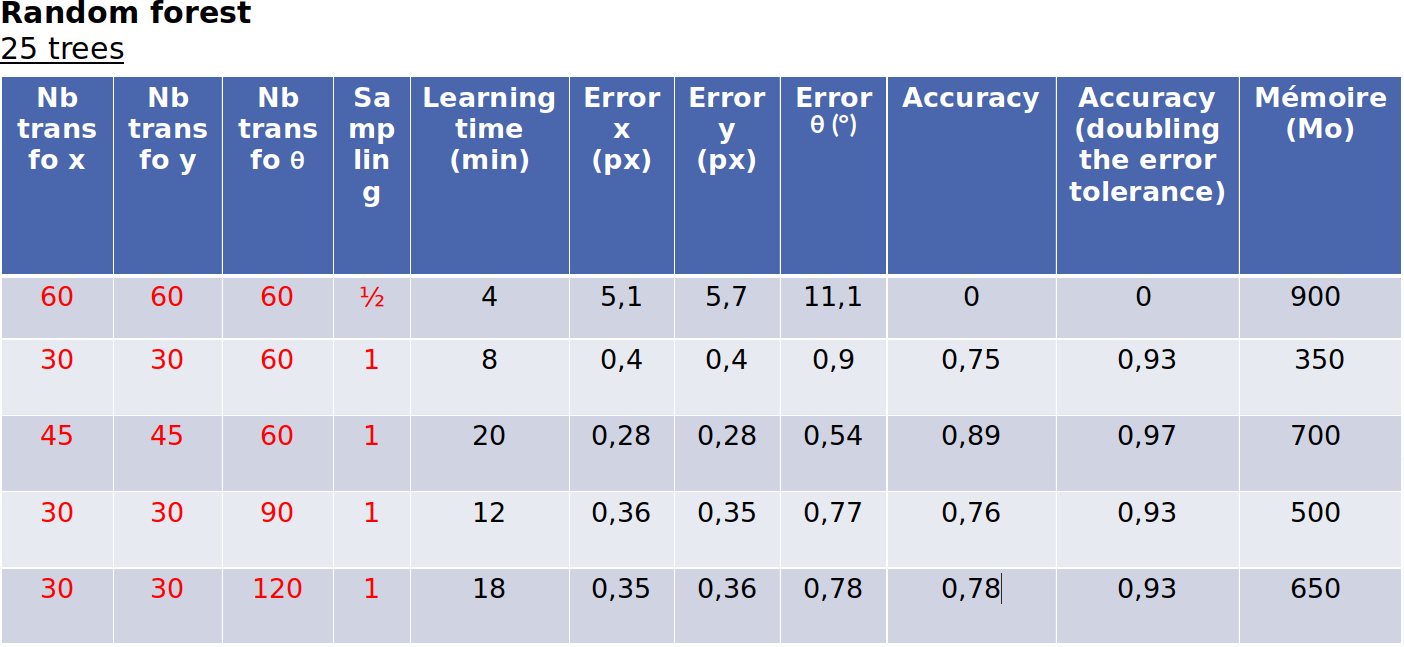
Here are some more testing with random forest 25 trees:
Another regression method tested was K-nearest neighbors. It has the advantage of being more robust to noise, having excellent prediction in both Atlas and acquisition dataset; 95% and a fast prediction time - 100ms.
Now with the atlas image the learning time is not much important as it would have to be trained only once, but again the memory required to store the model is too high, 1.5GB. In a 3D scenario it would again explode.
This concludes that it is not viable to determine the acquisition plans using Knn and random forest, when we consider all position and orientation constraints as unknowns.
Download the atlas image allenBrainAtlas.nii and the acquisition image CTRL_alc_M1_t2_1.nii.
Use delete_NaN() to remove NaN values of your image block.
Call create_dataset() passing the path to the one of the images downloaded.
Call get_fit() passing the dataset and labels to get the model.
Call predict() passing the model and a test image.
Call draw_plots() to visualize the accuracy and error curves.