4D (time evolving) depth S-grids#660
Conversation
…native to SWASH and allowing time varying depths in RectilinearSgrid for both defer and full load
for grid dim
parcels/field.py
Outdated
| kwargs['depth_field'] = dimensions['depth'][6:] | ||
| else: | ||
| depth = filebuffer.read_depth | ||
| print('Temporary print rather than error: Time varying depth data cannot be read in netcdf files yet') |
There was a problem hiding this comment.
Does this temporary print need to be here?
There was a problem hiding this comment.
Also, would this require a tutorial to explain to users how to use this new 4D S-grid functionality?
There was a problem hiding this comment.
happy to help creating one. I have plenty of (light) simulations that could be cleaned up and used for it.
parcels/field.py
Outdated
| return np.zeros(1) | ||
|
|
||
| @property | ||
| def read_depth_dim(self): |
There was a problem hiding this comment.
Confusingly named method? Would this be better read_depth_4d or so? And the other one read_depth_1d?
and the depth field is set with `set_depth_from_field`
|
Some small issue: |
|
I seem not authorised to merge the master into the origin/branch but locally I have merged since the update for the memory leak. There is still the same memory problem as discussed in #703 #711 and #668 and even with the master as it was this morning, I still seem to have it (in 1h I ran out of my RAM). It seemed solved and tested for other users so I was wondering @erikvansebille or @CKehl, could you merge the master in this branch as there are no conflicts (yet) and then if we're all on the same page we could hopefully fix where the memory issue still seems to persist. |
|
Hi @cjongedijk. I have just merged We have done (and are still doing) quite some optimisations of memory usage, so it may be that this updated version fixes your RAM issues. See also http://oceanparcels.org/faq.html#performance Let us know how it goes! |
|
Ho - so, I also encounter still some problems with long-running simulations, but not due to the memory but because there are file access conflicts, which I intent to resolve soon. Can you confirm (via the procedure in http://oceanparcels.org/faq.html#performance) that your error actually comes from memory overflow ? |
|
Also: I am right now encountering some issues with 4D data myself when compiling the MPI documentation notebook for 4D simulations, so we're working very much on the same issue, I believe. |
|
@erikvansebille thanks for the merge. @CKehl are you using this branch to simulate with 4D data? I haven't ran into any problems yet but @delandmeterp and me worked on it indeed but nothing has changd since I have not had any issues and could not really do anything or check any performance differences to the 3D fields with memory running out so fast on my longer sims. I should have flagged earlier that I cannot merge myself but I have locally been merging all the time and since I was understanding being the only user of this branch didnt bother pushing back to the repository. I use 4D fields for all of my simulations, however only the ones where the fields are large I run into memory problems, not for ones that have small fields and run for a longer time. I have monitored my memory over two simulations I did this week, one that has run (and is still running) with small fields and one that ran out of memory in one hour with large fields (similar pset sizes). I will post here in a minute how it looks and we can continue from there. |
|
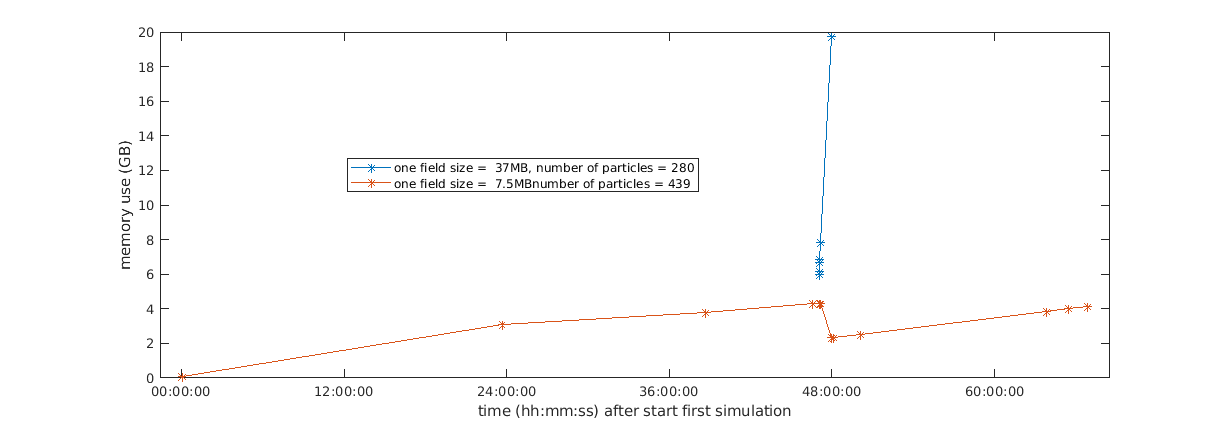
so this is how it looks like for me. I randomly exported the memory during the last few days sorry for the intermittent signal compared to the graphs @CKehl showed in the other threads , working on an automated memory tracker for python processes only but haven't done that yet. approximately 2 days in my slowly aggregating first (light) simulation, I had started one that gives me lots of troubles. I swapped after about 1 hour and then I killed the simulation. I had the long running one on purpose in the background because I saw there was a memory drop earlier whilst simultaneously running sims. This seems to indeed the case around 48 h into the sim where I kill the trouble simulation and the other one continues on a lower memory load. |
|
with 'one field size' I mean the size of the netcdf file containing all variables/field information for one time step. |
|
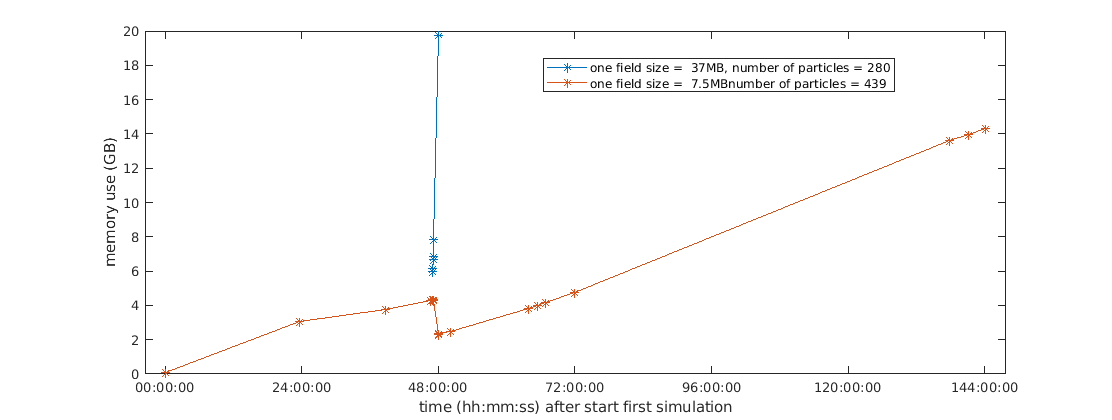
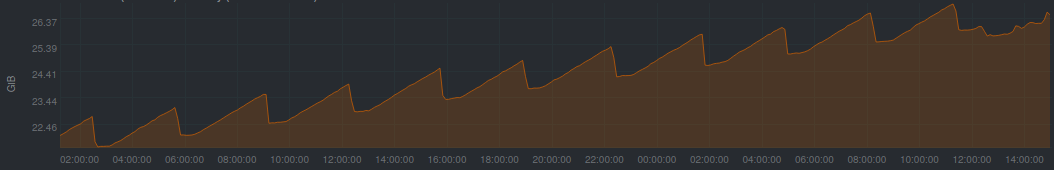
updated: after the weekend my 'light' simulation linearly increased the memory uptake as expected, and with netdata I tracked yesterday where my local system was only doing the parcels run with similar memory behaviour as in #703 https://github.com/OceanParcels/parcels/issues/668#issuecomment-576667920 tomorrow morning I'll have to kill it to free up space for other projects. Fig 1: Also, following the conversation in #740 today is that testing for _fromnemo specifically or also generally for _fromNetCDF? It seems like that data format of NEMO (sigma grids 3d+time, C grid) generally is very similar to mine. |
|
Hi Cleo - sorry to come back to you that late. Indeed this memory issue was keeping me up now the last 2 weeks, cause when I fixed it initially, I ran all tests just with 2D+time data. In 3D+time, I ran into massive issues because I wasn't converting the field_chunksize things properly for those cases - they are more 'involved'. Also, there was a separate issue popping up very often when running Erik's Galapagos case - conflicting file access, because with the lazy loading now actually implemented properly, the locking mechanism needs to be managed. As you already saw in #740 , those issue should be resolved now. I tested #740 by running the full galapagos case (the backward simulation) as it is in the repository (https://github.com/OceanParcels/GalapagosBasinPlastic) successfully on the cluster in a job system without crashes, using auto-chunking. Thus, if we can rebase this branch (update it to the current master), then it would be worth testing again. I am also running the python notebook (https://nbviewer.jupyter.org/github/OceanParcels/parcels/blob/master/parcels/examples/documentation_MPI.ipynb) now on a cluster with the 3D data, regenerating the images. It would be very kind if you were running you simulation with the updates again. Conversly, I'll now browse through your changes in this branch, make a review, and get some inspiration from your changes were I may still need to adapt things. |
|
BTW: if you wanna see how to define |
 CKehl
left a comment
CKehl
left a comment
There was a problem hiding this comment.
Looks good so far from the functionality - also didn't see anything that could conflict with other functionalities. Still, tests from @cjongedijk will be very important. That is because, if you work with a high-resolution depth column so that you actually prefer to also chunk the depth, then all the computattions asking for field.grid.depth[0] and field.grid.depth[-1] may either cause conflicts (but that's just a hypothesis).
Rename `depth_index` to `depth_indices` to highlight that it can be a vector
…aused the tests to fail.
…le data repository. Also requires a fix cause the import currently fails when trying to determine the minimal depth from the initial-empty grid(set).
… stuck with the varying-grid/field issue from #782.
…necessary check-size console prints
| dask.config.set({'array.chunk-size': '128MiB'}) | ||
| field_set = fieldset_from_swash(chunk_mode) | ||
| npart = 20 | ||
| # lonp = [i for i in np.arange(start=9.1, stop=11.3, step=0.1)[0:20]] |
| npart = 20 | ||
| # lonp = [i for i in np.arange(start=9.1, stop=11.3, step=0.1)[0:20]] | ||
| lonp = [i for i in 9.5 + (-0.2 + np.random.rand(npart) * 2.0 * 0.2)] | ||
| # latp = [i for i in 12.7+(-0.25+np.random.rand(npart)*2.0*0.25)] |
There was a problem hiding this comment.
another stray debug comment?
| if chunk_mode == 'auto': | ||
| chs = 'auto' | ||
| elif chunk_mode == 'specific': | ||
| # chs = {'x': 4, 'j': 4, 'z': 7, 'z_u': 6, 't': 1} |
|
|
||
| def fieldset_from_swash(chunk_mode): | ||
| filenames = path.join(path.join(path.dirname(__file__), 'SWASH_data'), 'field_*.nc') | ||
| # filenames = [] |
In this PR, we provide a
Field.from_netcdfthat allows to read vertical grids evolving in time. Thedepthdimension must beField.depthfieldwhere 'depthfieldis the name of the field on which the data of the depth are provided, such thatU.grid.depth= depthfield.data`