Conversation
|
@tdunning Please take a look at the modified test and see if limiting the error to be within 2% make sense for compression 200? |
Codecov Report
@@ Coverage Diff @@
## master #7076 +/- ##
============================================
- Coverage 73.64% 73.57% -0.07%
Complexity 91 91
============================================
Files 1480 1480
Lines 72871 72871
Branches 10483 10483
============================================
- Hits 53664 53614 -50
- Misses 15734 15781 +47
- Partials 3473 3476 +3
Flags with carried forward coverage won't be shown. Click here to find out more.
Continue to review full report at Codecov.
|
I can't comment exactly without understanding the test. I can see that 200,000 random numbers are generated and put pair-by-pair into t-digests that are somehow serialized and combined. Ultimately, the data seems to be grouped using two fields each with 100 values resulting in a total 10,000 digests each with an average of 20 data points in them. Beyond this outline, my understanding fades to nothing. Can you say what the different between the star-index path and non-star-index path is? Without understanding the details, I think that it is a reasonable assumption to say that the difference between two digests with the same data but constructed differently will be limited by the difference between difference between either digest and the actual empirical distribution. The test, as designed, does not appear to be likely to put more than 45 data points in a single digest which means that any compression parameter greater than 100 will result in zero error because all 45 data points will be preserved. My guess is that the underlying intent of the test is a bit different from the execution, though, and you are interested in two things over a range of digest sizes:
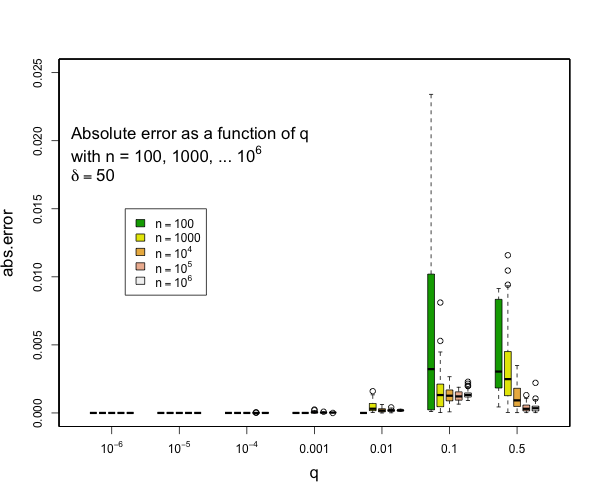
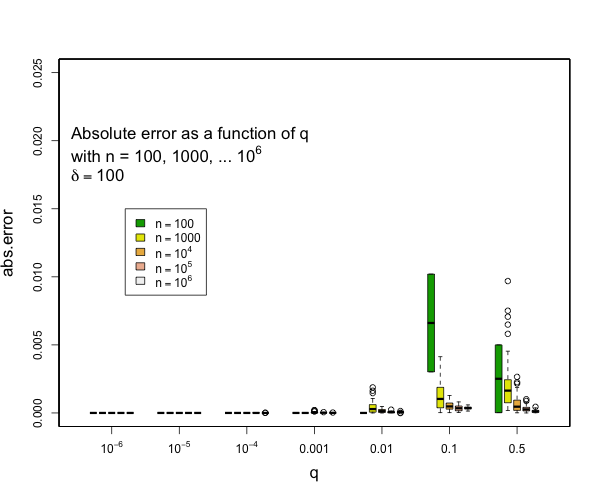
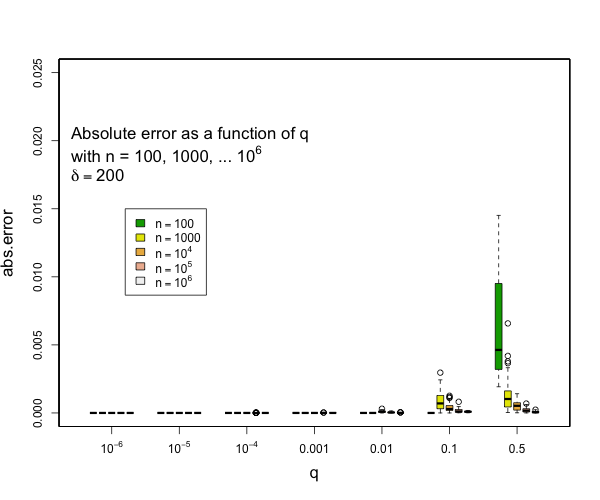
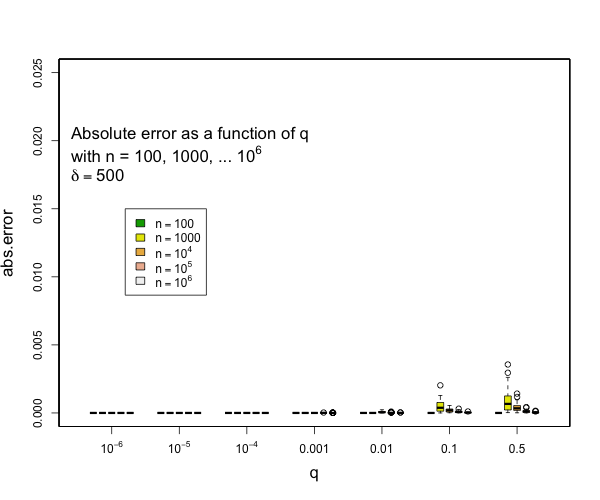
I can say that pretty much all quantiles such that q <= 0.1 or q >= 0.9 will have really small errors with compression of 200 for as many samples as you might expect and still quite small errors with compression of 100. Here are some examples derived from 50 runs for different values of compression (delta). In general, the worst case is when there is a transition to two samples per centroid. Other than this, the accuracy for the median is generally worse than the accuracy for the tails (that is, of course, the point of the t-digest). In general, for 100 samples or more, you can occasionally have errors of more than 2% with delta=50, and will rarely have errors over 1% with delta=100 or delta=200. I don't have handy data for n < 100, for delta=200, there should never be any errors for such small counts except possibly right at the median. |
|
@tdunning Thanks for the explanation. |
|
It is not expected that accuracy would be worse if you normalize by number of centroids. On the other hand, compression 200 with new should by at least as many centroids as compression 100 in 3.2. Can you say more about the merge strategies of interest? I would be happy to build a reference test that mirrors your needs. Merging through a ser-de step will definitely have an impact on accuracy since more data is transiently kept in memory than is strictly specified by the compression level. Even so, we should be able to find a good trade-off. |
|
The random t-digest objects are created as following: In non-star-tree approach simply merges the t-digests in sequence without ser-de; the star-tree approach will pre-aggregate t-digests and then stores the serialized merged t-digests (each pre-aggregated t-digest might go through multiple rounds of ser-de). Non-star-tree: Star-tree: Then we compare the result for each quantile from these 2 result t-digest objects. Hope this can explain the test logic. |
|
Yes. Very helpful.
I have a test that does roughly this. I keep the raw data for comparison, a
t-digest that is never serialized, one that is serialized and deserialized
for each sample and five that each get random subsets of data and then are
merged at the end. The final analysis looks at discrepancies against exact
answers and the three digested approaches. Not done yet.
…On Mon, Jun 21, 2021 at 11:28 AM Xiaotian (Jackie) Jiang < ***@***.***> wrote:
@tdunning <https://github.com/tdunning>
The random t-digest objects are created as following:
@OverRide
Object getRandomRawValue(Random random) {
TDigest tDigest = TDigest.createMergingDigest(COMPRESSION);
tDigest.add(random.nextInt(MAX_VALUE));
tDigest.add(random.nextInt(MAX_VALUE));
return ObjectSerDeUtils.TDIGEST_SER_DE.serialize(tDigest);
}
In non-star-tree approach simply merges the t-digests in sequence without
ser-de; the star-tree approach will pre-aggregate t-digests and then stores
the serialized merged t-digests (each pre-aggregated t-digest might go
through multiple rounds of ser-de).
Non-star-tree:
TDigest tDigest = deserialize(tDigest1);
tDigest.add(deserialize(tDigest2));
tDigest.add(deserialize(tDigest3));
tDigest.add(deserialize(tDigest4));
Star-tree:
TDigest tDigest = deserialize(tDigest1);
tDigest.add(deserialize(tDigest2));
byte[] mergedTDigest1 = serialize(tDigest);
tDigest = deserialize(tDigest3);
tDigest.add(deserialize(tDigest4));
byte[] mergedTDigest2 = serialize(tDigest);
tDigest = deserialize(mergedTDigest1);
tDigest.add(deserialize(mergedTDigest2));
Then we compare the result for each quantile from these 2 result t-digest
objects.
Hope this can explain the test logic.
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
<#7076 (comment)>,
or unsubscribe
<https://github.com/notifications/unsubscribe-auth/AAB5E6XDZ2GJYY3ZE5JDYL3TT6AE7ANCNFSM47AL7QEA>
.
|
|
@tdunning Any update on this? |
|
Yes. There is an update.
I think that there is a fundamental problem with the way that the test is
phrased.
Any digest that has limited memory will ultimately have to lose some
information and will estimate quantiles. That will lead to errors in sample
space roughly equal to the separation between adjacent samples. Different
digests may keep more samples or fewer and thus adjust when this happens
and they may prioritize samples in different parts of the distribution, but
there will always be an approximation error when samples are combined.
This means that if the samples are more than 0.02 x max apart, you have the
risk of error unless the digest is not a digest at all. The max separation
for N uniform random samples has a 99% percentile value of 4.6 / N. That
means that you will need 230 samples before you have a 99% chance of
having no gap bigger than 99%. In turn, that means that your digest will
need to keep at least 230 samples before coalescing to be reasonably
assured of passing the test. Some digests might do a bit better than this,
but not massively. For instance, the t-digest will focus coalescence into a
fraction of the interval and thus have a probability of failure of 1/2 to
1/10 of this value.
On the other hand, if you recast the error back into quantile space by
calculating the empirical CDF of the estimated quantile, you immediately
have a well-behaved system. Your unavoidable errors will be bounded by 1/N
and a t-digest that keeps 50 samples or more and limits errors to one
sample size should have the guarantees that you want.
…On Thu, Jul 1, 2021 at 10:42 AM Xiaotian (Jackie) Jiang < ***@***.***> wrote:
@tdunning <https://github.com/tdunning> Any update on this?
—
You are receiving this because you were mentioned.
Reply to this email directly, view it on GitHub
<#7076 (comment)>,
or unsubscribe
<https://github.com/notifications/unsubscribe-auth/AAB5E6VIUKHHS5FQAXWXWEDTVSSKDANCNFSM47AL7QEA>
.
|
|
@tdunning Thanks for the explanation. |
|
@richardstartin Do you want to take a look at this discussion thread? We observed worse accuracy after upgrading the library, but not sure if that is expected because of a more advanced algorithm (e.g. tradeoff between space and accuracy) |
Also modify the test per the discussion in #7069