[eda] added explain_rows method to autogluon.eda.auto - Kernel SHAP visualization#3014
Conversation
explain_rows method to autogluon.eda.auto; the methods performs Kernel SHAP values analysis and visualizationexplain_rows method to autogluon.eda.auto - Kernel SHAP visualization
| 'phik>=0.12.2,<0.13', | ||
| 'seaborn>=0.12.0,<0.13', | ||
| 'ipywidgets>=7.7.1,<9.0', # min versions guidance: 7.7.1 collab/kaggle | ||
| 'shap>=0.41,<0.42', |
There was a problem hiding this comment.
@gradientsky FYI, fastshap may be of interest to test out and compare performance as suggested here: #2222 (comment)
fastshap claims to be much faster than shap:
There was a problem hiding this comment.
Did a code rework. The code now split into backend analysis and rendering parts. Analysis supports both shap and fastshap libraries (two different backends). Using shap as a default one for auto functionality because it is faster.
Visualizations also split into separate primitives; compatible with both of the backends.
|
Job PR-3014-c5e116b is done. |
c5e116b to
4c93e16
Compare
|
Check out this pull request on See visual diffs & provide feedback on Jupyter Notebooks. Powered by ReviewNB |
4c93e16 to
f1229f8
Compare
|
Blocked by AnotherSamWilson/fastshap#8 |
eda/setup.py
Outdated
| 'seaborn>=0.12.0,<0.13', | ||
| 'ipywidgets>=7.7.1,<9.0', # min versions guidance: 7.7.1 collab/kaggle | ||
| 'shap>=0.41,<0.42', | ||
| 'fastshap>=0.3,<0.4', |
There was a problem hiding this comment.
Is fastshap good enough to have as a required dependency? What is the relative advantages of fastshap over shap?
There was a problem hiding this comment.
For the purposes of explaining a few rows here, fastshap is slower than shap. I sent an update to remove fastshap backend completely (performance + support concern).
…SHAP values analysis and visualization
4a6d17d to
2bc659e
Compare
|
Job PR-3014-4a6d17d is done. |
|
Job PR-3014-2bc659e is done. |
|
Job PR-3014-c4fc440 is done. |
 Innixma
left a comment
Innixma
left a comment
There was a problem hiding this comment.
LGTM! Had some minor comments
| def predict_proba(self, X): | ||
| if isinstance(X, pd.Series): | ||
| X = X.values.reshape(1, -1) | ||
| if not isinstance(X, pd.DataFrame): | ||
| X = pd.DataFrame(X, columns=self.feature_names) | ||
| if self.ag_model.problem_type == REGRESSION: | ||
| preds = self.ag_model.predict(X) | ||
| else: | ||
| preds = self.ag_model.predict_proba(X) | ||
| if self.ag_model.problem_type == REGRESSION or self.target_class is None: | ||
| return preds | ||
| else: | ||
| return preds[self.target_class] |
There was a problem hiding this comment.
I see a similar hack here that I see in the general AutoGluon Tabular code of using predict_proba for regression. Long term do you think this is the right thing to do long-term? is it used to simplify the code logic or just to align with how Tabular does things?
| logger = logging.getLogger(__name__) | ||
|
|
||
|
|
||
| class _ShapAutogluonWrapper: |
There was a problem hiding this comment.
_ShapAutogluonWrapper or _ShapAutoGluonWrapper?
| baseline_sample: int, default = 100 | ||
| The background dataset size to use for integrating out features. To determine the impact | ||
| of a feature, that feature is set to "missing" and the change in the model output | ||
| is observed. |
There was a problem hiding this comment.
any guidance on what magnitudes represent what noise level?
Is 100 enough? 1000? Will 10000 differ significantly from 100?
Why choose 100 when I could choose 10000? Is it purely to save compute time?
Will it still work if I set it to 1?
There was a problem hiding this comment.
"There should be enough rows to cancel-out significant variance in the base value"
Do we assume the user knows what "enough rows" is? Would it be better to give guidance in the doc-string?
| rows to explain | ||
| baseline_sample: int, default = 100 | ||
| The background dataset size to use for integrating out features. To determine the impact | ||
| of a feature, that feature is set to "missing" and the change in the model output |
There was a problem hiding this comment.
"missing": Do we have a consistent definition for what setting to "missing" means?
There was a problem hiding this comment.
This is how the shap describes the process.
| else: | ||
| _baseline_sample = len(args.train_data) | ||
|
|
||
| baseline = args.train_data.sample(_baseline_sample, random_state=0) |
There was a problem hiding this comment.
move random_state=0 to an init arg?
| for _, row in self.rows.iterrows(): | ||
| _row = pd.DataFrame([row]) | ||
| if args.model.problem_type == REGRESSION: | ||
| predicted_class = 0 |
There was a problem hiding this comment.
Good call; updated
|
|
||
| misclassified = y_proba[y_true_val != y_pred_val] | ||
| expected_value = misclassified.join(y_true_val).apply(lambda row: row.loc[row[label]], axis=1) | ||
| predicted_value = misclassified.max(axis=1) |
There was a problem hiding this comment.
Long-term we may need to revisit this, as .max might not actually be identical to the predicted value. For example, if we start maximizing f1 score, we would set a threshold that isn't 0.5. Probably not necessary to address in this PR, but maybe worth adding a TODO.
There was a problem hiding this comment.
I'd imagine we eventually make a method like pred = predictor.proba_to_pred(proba, metric='f1') which contains an inner dictionary of metric -> threshold such as {'f1': 0.63} that influences the pred that is returned
There was a problem hiding this comment.
Nevermind this specific comment, I see predictor.predict called above, so it should be fine.
| baseline_sample: int, default = 100 | ||
| The background dataset size to use for integrating out features. To determine the impact | ||
| of a feature, that feature is set to "missing" and the change in the model output | ||
| is observed. |
There was a problem hiding this comment.
Very random thought completely tangential to the PR: I wish IDEs/code had logic where we could specify a docstring for a parameter one time in some file, then have it auto-populate via a variable reference in the source-code docs so we just have to write
baseline_sample: int, default = 100
{baseline_sample_docstring}
and the IDE magically converts it to text unless we click to force it to show source, so we avoid having to copy/paste the same doc-string many times across many usages.
That is my random thought of the day. Carry on.
|
Job PR-3014-5c757a8 is done. |
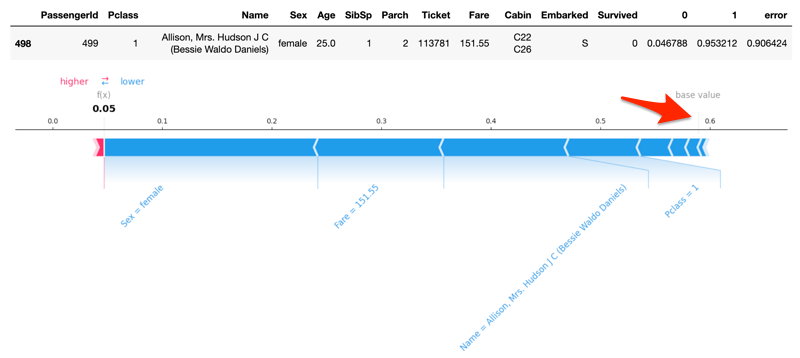
Description of changes:
explain_rowsmethod toautogluon.eda.auto; the methods performs Kernel SHAP values analysis and visualizationquick_fit: fixes for highest_error and undecided rows calculationsExamples
Using primitives
By submitting this pull request, I confirm that you can use, modify, copy, and redistribute this contribution, under the terms of your choice.