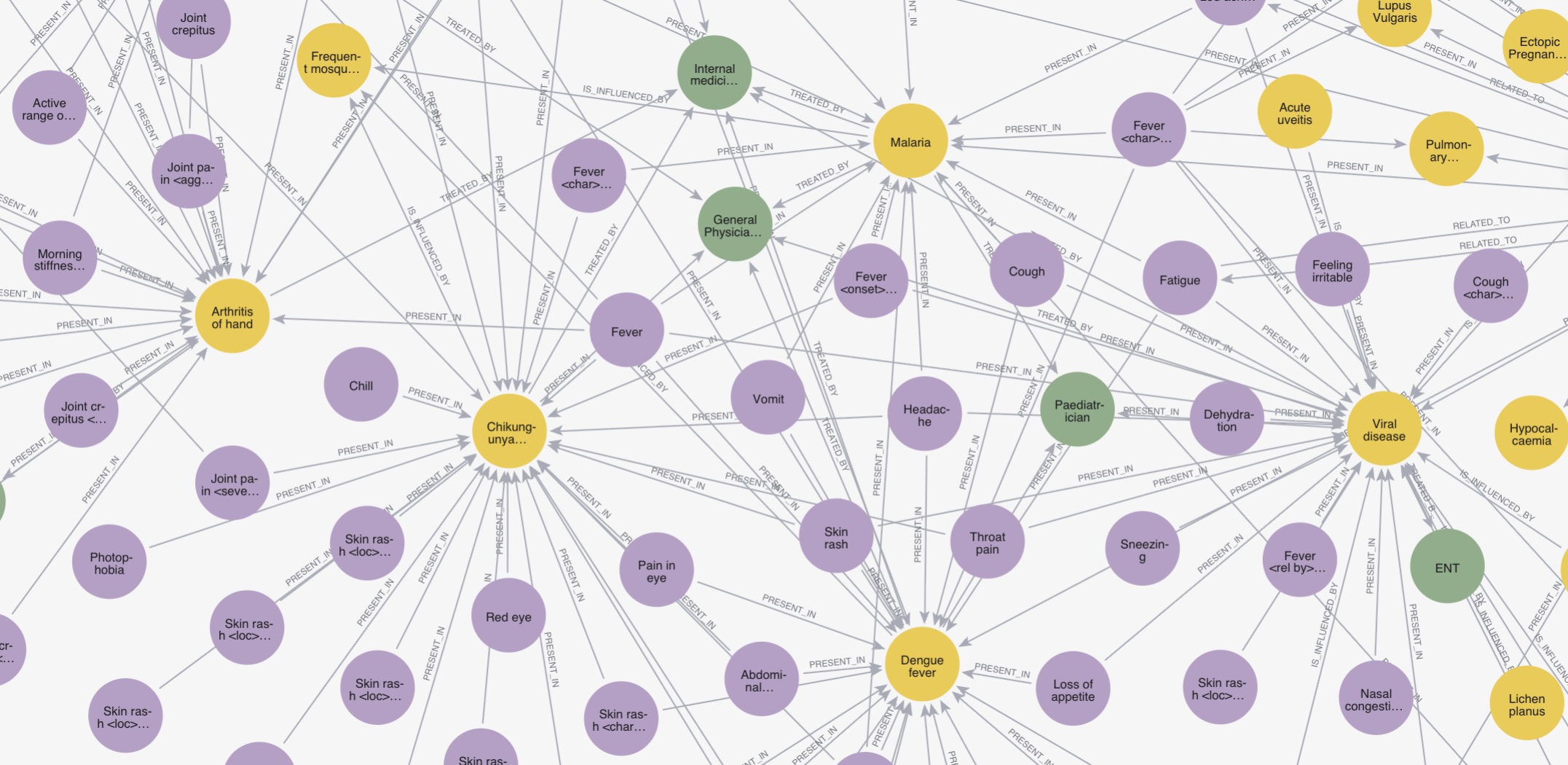
Open clinical knowledge graphs for grounding healthcare AI
SNOMED-linked knowledge graphs built to ground healthcare AI in verified clinical facts. Developed at Eka Care and battle-tested in production across symptom checking, differential diagnosis, and patient health profiling. Released openly under CC BY-NC 4.0.
→ Read the full writeup on the motivation, design, and use cases
| Network | Focus | Nodes | Relationships |
|---|---|---|---|
| bodhi-s | Condition ↔ Symptom ↔ Speciality | 4,855 | 13,204 |
| bodhi-m | Concept ↔ Drug ↔ LabInvestigation | 4,469 | 3,566 |
Each network is available in five formats:
| Format | File | Use case |
|---|---|---|
| Neo4j dump | neo4j/*.dump |
Import into a local Neo4j instance |
| CSV | csv/*.csv |
Flat-file processing, Pandas, SQL |
| JSONL | jsonl/triples.jsonl, jsonl/nl_facts.jsonl |
LLM training, RAG pipelines |
| PyG | pyg/*.pt |
PyTorch Geometric / GNN training |
| RDF/Turtle | rdf/*.ttl |
Semantic web, SPARQL, ontology tools |
| Browser JSON | browser_data_*.json |
Graph visualisation (nodes + edges in one file) |
# Restore into a local Neo4j instance (Neo4j 5.x)
neo4j-admin database load --from-path=bodhi-s/neo4j/bodhi.dump --database=neo4j --overwrite-destination=trueimport pandas as pd
nodes_condition = pd.read_csv("bodhi-s/csv/nodes_condition.csv")
edges_present_in = pd.read_csv("bodhi-s/csv/edges_present_in.csv")import torch
from torch_geometric.data import HeteroData
data = torch.load("bodhi-s/pyg/bodhi.pt")
print(data)# Using Apache Jena's riot / arq
arq --data bodhi-s/rdf/bodhi.ttl --query your_query.sparqlLinks conditions to symptoms, specialities, and inter-condition risk relationships. Symptoms are modelled as compound variants (e.g. Fever with chills, Fever for 3 days) with per-node triage levels and demographic likelihood scores.
| Metric | Count |
|---|---|
| Condition nodes | 779 |
| Symptom nodes (variants) | 4,037 |
| Symptom root concepts (distinct SNOMED IDs) | 590 |
| Speciality nodes | 39 |
| Total relationships | 13,204 |
| Symptom → Condition edges (PRESENT_IN) | 10,352 |
| Condition → Speciality edges (TREATED_BY) | 1,558 |
| Condition → Condition edges (IS_INFLUENCED_BY) | 1,020 |
| Condition → Condition edges (RELATED_TO) | 221 |
| Condition → Condition edges (HAS_PREREQUISITE) | 53 |
Condition types: Disorder 607 · Misc 84 · FamilyHistory 49 · Lifestyle 21 · Procedure 16 · Allergy 1 · Symptom 1
Avg symptoms per condition: 13.3
Condition triage: OPD Managed 367 (47%) · Worrisome 223 (29%) · Emergency 189 (24%)
Symptom triage: OPD Managed 2,244 (56%) · Worrisome 1,540 (38%) · Emergency 252 (6%)
Most cross-cutting symptoms: Fever (145 conditions) · Fatigue (126) · Headache (110) · Vomit (94) · Malaise (81)
Top specialities by condition volume: Internal Medicine (292) · General Physician (205) · Orthopedic (139) · Neurologist (83) · General Surgeon (81)
A diagnosable medical condition, disorder, or clinical entity.
| Property | Type | Values | Description |
|---|---|---|---|
snomed_id |
string | SNOMED CT ID | Globally unique clinical identifier |
name |
string | — | Clinical name of the condition |
concept_type |
enum | Disorder Misc FamilyHistory Lifestyle Procedure Allergy Symptom |
Classification of the concept |
triage_level |
enum | opd_managed worrisome emergency |
Clinical urgency |
type_condition |
enum | acute chronic acute_that_may_turn_chronic chronic_with_acute_aggravation lifestyle medical_history Event Injury |
Temporal nature of condition |
overall_likelihood |
enum | rare low medium high very_high |
Population prevalence signal |
likelihood_male |
float | 0.0–1.0 | Relative likelihood in males |
likelihood_female |
float | 0.0–1.0 | Relative likelihood in females |
likelihood_age_0_1 |
float | 0.0–1.0 | Relative likelihood in age 0–1 |
likelihood_age_1_5 |
float | 0.0–1.0 | Relative likelihood in age 1–5 |
likelihood_age_6_12 |
float | 0.0–1.0 | Relative likelihood in age 6–12 |
likelihood_age_13_18 |
float | 0.0–1.0 | Relative likelihood in age 13–18 |
likelihood_age_19_30 |
float | 0.0–1.0 | Relative likelihood in age 19–30 |
likelihood_age_30_45 |
float | 0.0–1.0 | Relative likelihood in age 30–45 |
likelihood_age_45_60 |
float | 0.0–1.0 | Relative likelihood in age 45–60 |
likelihood_age_60_plus |
float | 0.0–1.0 | Relative likelihood in age 60+ |
A clinical symptom or refinement thereof. Symptoms are structured with parent-child refinements (e.g. "Headache > Throbbing headache > Throbbing headache on right side").
| Property | Type | Values | Description |
|---|---|---|---|
uuid |
string | UUID | Unique compound symptom identifier |
snomed_id |
string | SNOMED CT ID | SNOMED identifier for the symptom |
root_snomed_id |
string | SNOMED CT ID | Parent/root symptom SNOMED ID |
root_snomed_name |
string | — | Parent symptom name |
name |
string | — | Full compound symptom name |
triage_level |
enum | opd_managed worrisome emergency |
Clinical urgency of this symptom |
relation1_type |
enum | characteristic severity location laterality onset duration_since duration_lasts temporal_pattern pain_type radiating aggravated relieved |
Type of the first refinement axis |
child1_name |
string | — | Value of the first refinement |
grouping1_selection_type |
enum | s m |
Single (s) or multi-select (m) for axis 1 |
relation2_type |
enum | (same as relation1_type) | Type of the second refinement axis |
child2_name |
string | — | Value of the second refinement |
grouping2_selection_type |
enum | s m |
Single or multi-select for axis 2 |
relation3_type |
enum | (same as relation1_type) | Type of the third refinement axis |
child3_name |
string | — | Value of the third refinement |
grouping3_selection_type |
enum | s m |
Single or multi-select for axis 3 |
A medical speciality or care discipline.
| Property | Type | Description |
|---|---|---|
id |
string | Internal speciality identifier |
name |
string | Speciality name e.g. Cardiologist |
Links a symptom to a condition it presents in. Encodes bidirectional likelihood.
| Property | Values | Description |
|---|---|---|
likelihood_symptom_given_condition |
zero rare low medium high very_high |
How commonly this symptom appears when the condition is present — P(symptom | condition) |
likelihood_condition_given_symptom |
zero rare low medium high very_high |
How predictive this symptom is of the condition — P(condition | symptom) |
Indicates which speciality manages a condition.
| Property | Values | Description |
|---|---|---|
weight |
rare low medium high very_high |
Strength of the referral association |
Condition A is clinically influenced by the presence of condition B (e.g. Diabetes IS_INFLUENCED_BY Obesity).
| Property | Values | Description |
|---|---|---|
relation_strength |
zero rare low medium high very_high |
Magnitude of influence |
relation_polarity |
positive negative |
Positive = B increases risk of A; Negative = B decreases risk of A |
Condition A requires condition B to be present (e.g. Diabetic nephropathy HAS_PREREQUISITE Diabetes mellitus).
| Property | Values | Description |
|---|---|---|
relation_strength |
medium high very_high |
How mandatory the prerequisite is |
relation_polarity |
positive negative |
Direction of dependency |
Ontological relatedness — used for symptom deduplication and SNOMED hierarchy linkage.
| Property | Values | Description |
|---|---|---|
relation_type |
same_as similar_to parent_child |
Nature of the ontological relationship |
Maps SNOMED concepts (disorders, findings, procedures) to generic drugs and LOINC-coded lab investigations, organised in a three-level hierarchy (System → Group → Granular). Supports reverse inference from medications to conditions, and from lab results to health domains.
| Metric | Count |
|---|---|
| Concept nodes | 2,471 |
| Drug nodes | 1,186 |
| LabInvestigation nodes | 812 |
| Total relationships | 3,566 |
| Concept → Concept edges (CHILD_OF) | 1,768 |
| Concept → Drug edges (TREATED_BY) | 908 |
| LabInvestigation → Concept edges (IMPACTS) | 808 |
| Concept → LabInvestigation edges (MONITORED_BY) | 82 |
Concept hierarchy: System 14 → Group 250 → Granular 1,942 (+ 265 unmapped)
Hierarchy coverage: 1,540 / 1,942 granular linked to group · 228 / 250 groups linked to system
A clinical concept — condition, disorder, finding, procedure, or lifestyle factor — organised in a three-level hierarchy: System → Group → Granular.
| Property | Type | Values | Description |
|---|---|---|---|
snomed_id |
string | SNOMED CT ID | Primary unique identifier (open standard) |
name |
string | — | Clinical name |
display_name |
string | — | Consumer-friendly name e.g. High Blood Pressure |
level_concept |
enum | system group granular |
Position in the clinical hierarchy |
type_concept |
enum | Disorder Finding Procedure Lifestyle Allergy Situation |
Clinical category |
type_information |
enum | SelfHistory FamilyHistory |
Whether this pertains to the patient themselves or family history |
active |
string | 1 |
Whether this concept is active in the knowledge base |
Hierarchy levels:
system— broad health domain e.g. Cardiovascular health, Endocrine health (14 nodes)group— clinical cluster e.g. Diabetes mellitus, Coronary Artery Disease (250 nodes)granular— specific diagnosable entity e.g. Diabetes mellitus type II (1,942 nodes)
A generic drug formulation. Combination drugs are stored as a single node.
| Property | Type | Description |
|---|---|---|
hash |
string | MD5 hash of the generic name — unique deduplication key |
name |
string | Generic drug name e.g. metformin, atorvastatin + aspirin |
therapeutic_class |
string | Comma-separated therapeutic class(es) e.g. Anti Diabetic, Cardiovascular |
A lab test or clinical measurement, identified by LOINC standard code.
| Property | Type | Values | Description |
|---|---|---|---|
loinc_id |
string | LOINC ID | Globally unique lab test identifier (open standard) |
name |
string | — | Standard test name |
display_name |
string | — | Display/friendly name |
system_map |
string | — | Health domain e.g. Renal health, Endocrine health |
timespan_problem |
enum | stat less_than_24_hr week_1 month_1 month_3 month_6 year_1 lifetime |
How long this lab investigation result stays clinically relevant for a related condition |
impact_problem |
enum | zero low medium high |
Clinical significance of this lab investigation in disease management |
Encodes the three-level clinical hierarchy. Granular concepts point to their parent Group; Group concepts point to their parent System.
No properties.
A lab investigation broadly belongs to and impacts a health domain concept (always a system-level concept). Represents the primary health system this test monitors.
No properties.
A clinical concept is treated by a generic drug. Encodes gender exclusivity signals for prescribing guidance.
| Property | Values | Description |
|---|---|---|
therapeutic_class |
string | The drug class relevant for this specific indication |
exclusivity |
zero low high |
How specific this drug is to this condition vs. used broadly |
exclusivity_male |
low high |
Prescribing exclusivity signal for males |
exclusivity_female |
low high |
Prescribing exclusivity signal for females |
A condition is monitored or diagnostically associated with a specific lab test. Encodes the deduction power and directional threshold.
| Property | Values | Description |
|---|---|---|
polarity |
above below equal |
Which direction relative to the threshold is clinically significant |
category_threshold |
normal borderline_low borderline_high high abnormal critically_high |
The result range that triggers this association |
vital_expiry_value |
stat week_1 month_1 month_3 month_6 year_1 |
The validity or the relevance of a test prescribed. Duration for which the vitals results are still considered relevant for a condition |
exclusivity |
low high |
Deduction power — how strongly an abnormal result predicts this condition |
BODHI/
├── bodhi-s/
│ ├── csv/ # Node and edge CSV files
│ │ ├── nodes_condition.csv
│ │ ├── nodes_symptom.csv
│ │ ├── nodes_speciality.csv
│ │ ├── edges_present_in.csv
│ │ ├── edges_treated_by.csv
│ │ ├── edges_is_influenced_by.csv
│ │ ├── edges_related_to.csv
│ │ └── edges_has_prerequisite.csv
│ ├── jsonl/
│ │ ├── triples.jsonl # (subject, predicate, object) triples
│ │ └── nl_facts.jsonl # Natural-language fact strings
│ ├── neo4j/
│ │ └── bodhi.dump # Neo4j database dump
│ ├── pyg/
│ │ └── bodhi.pt # PyTorch Geometric HeteroData object
│ ├── rdf/
│ │ └── bodhi.ttl # RDF graph in Turtle format
│ └── browser_data_bodhi_s.json # Combined nodes + edges for visualisation
└── bodhi-m/
├── csv/
│ ├── nodes_concept.csv
│ ├── nodes_drug.csv
│ ├── nodes_lab_investigation.csv
│ ├── edges_child_of.csv
│ ├── edges_treated_by.csv
│ ├── edges_impacts.csv
│ └── edges_monitored_by.csv
├── jsonl/
│ ├── triples.jsonl
│ └── nl_facts.jsonl
├── neo4j/
│ └── bodhi_m.dump
├── pyg/
│ └── bodhi_m.pt
├── rdf/
│ └── bodhi_m.ttl
└── browser_data_bodhi_m.json
| Standard | Used for |
|---|---|
| SNOMED CT | Condition and concept identifiers |
| LOINC | Lab investigation identifiers |
This dataset is released under the Creative Commons Attribution-NonCommercial 4.0 International (CC BY-NC 4.0) license.
You are free to share and adapt the data for non-commercial purposes, provided you give appropriate credit to Eka Care.