Probably a question more than an issue...
When I run pytest -v, one check fails. Below is the error message.
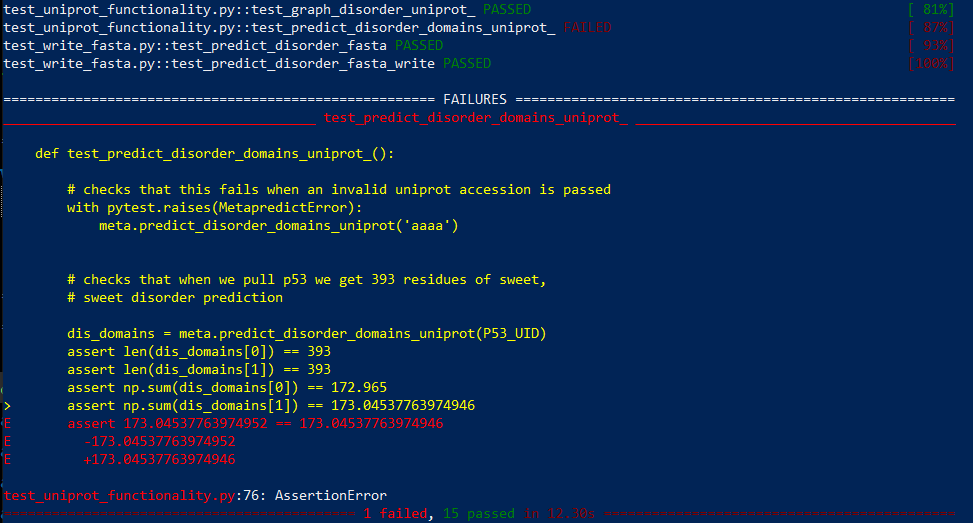
I'm running from a Windows machine with conda: do you think this is caused by the known clash with conda-installed numpy, or is this simply a rounding difference? When I run the test manually within my own python script on a manually downloaded p53 sequence from Uniprot and use np.sum() for the test, I also get the same value that occurred during the pytest (173.04537763974952). However, when I run the same test with Python's built-in sum() function, I get 173.04537763974955 (maybe this also is a known behavior difference between np.sum() and built-in sum()).
More importantly, given the miniscule difference between the values compared by pytest, is it safe to say that identical regions are still being classified as disordered?
Thanks!
-Sean
Probably a question more than an issue...

When I run pytest -v, one check fails. Below is the error message.
I'm running from a Windows machine with conda: do you think this is caused by the known clash with conda-installed numpy, or is this simply a rounding difference? When I run the test manually within my own python script on a manually downloaded p53 sequence from Uniprot and use np.sum() for the test, I also get the same value that occurred during the pytest (173.04537763974952). However, when I run the same test with Python's built-in sum() function, I get 173.04537763974955 (maybe this also is a known behavior difference between np.sum() and built-in sum()).
More importantly, given the miniscule difference between the values compared by pytest, is it safe to say that identical regions are still being classified as disordered?
Thanks!
-Sean