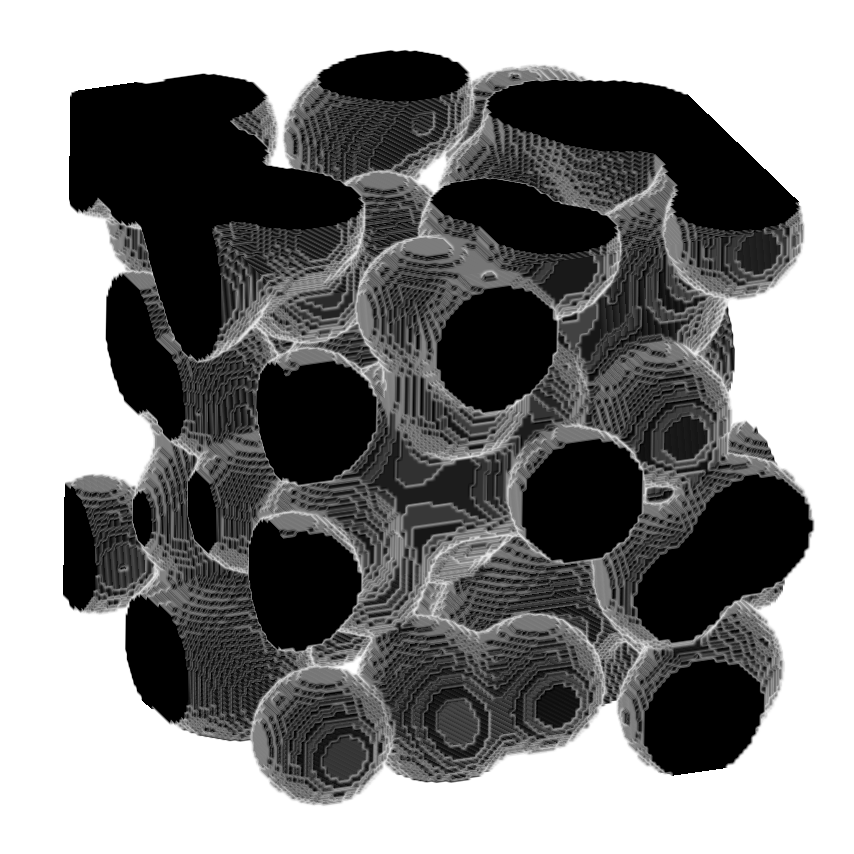
This package extends the scikit-image function medial_axis to the 3D case.
pip install medialaxis3dAutomatically installed with pip:
numpyscipycython
Optional only for visualization
napari
WIP
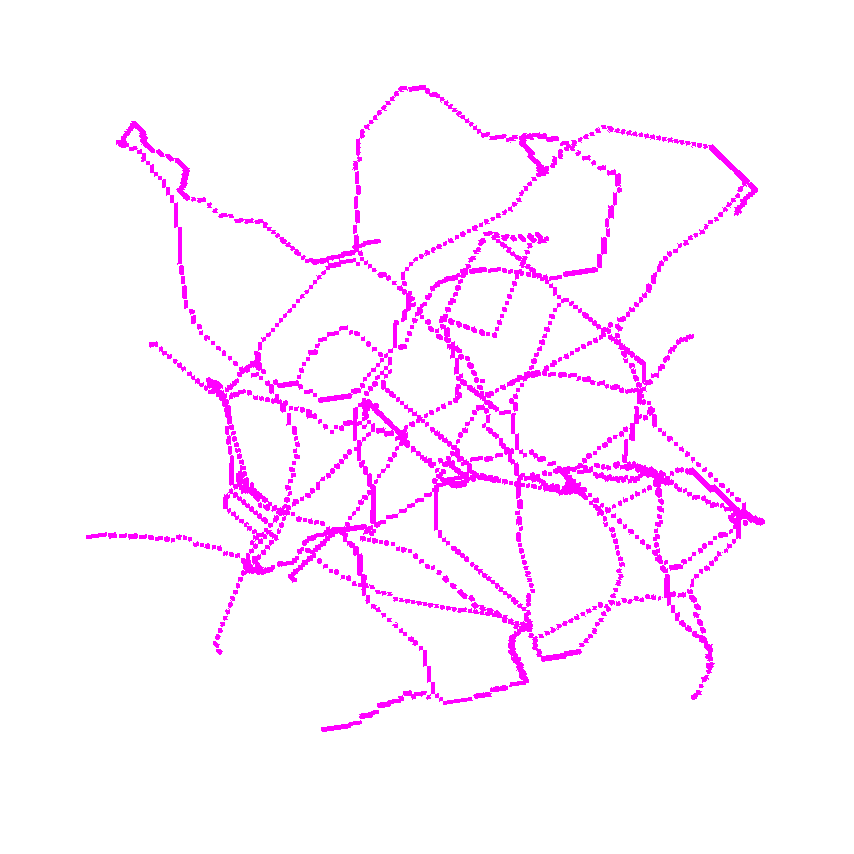
Use it without returning the medial distance.
>>> import numpy as np
>>> import skimage as ski
>>> import medialaxis3d
>>> import napari
>>> rng = np.random.default_rng(1278)
>>> image = ski.data.binary_blobs(length = 128,
>>> blob_size_fraction = 0.2,
>>> n_dim = 3,
>>> volume_fraction = 0.6,
>>> rng = rng)
>>> skeleton = medialaxis3d.medial_axis_3d(image,
>>> return_distance = False,
>>> size = 8,
>>> rng = rng)
>>> viewer = napari.Viewer()
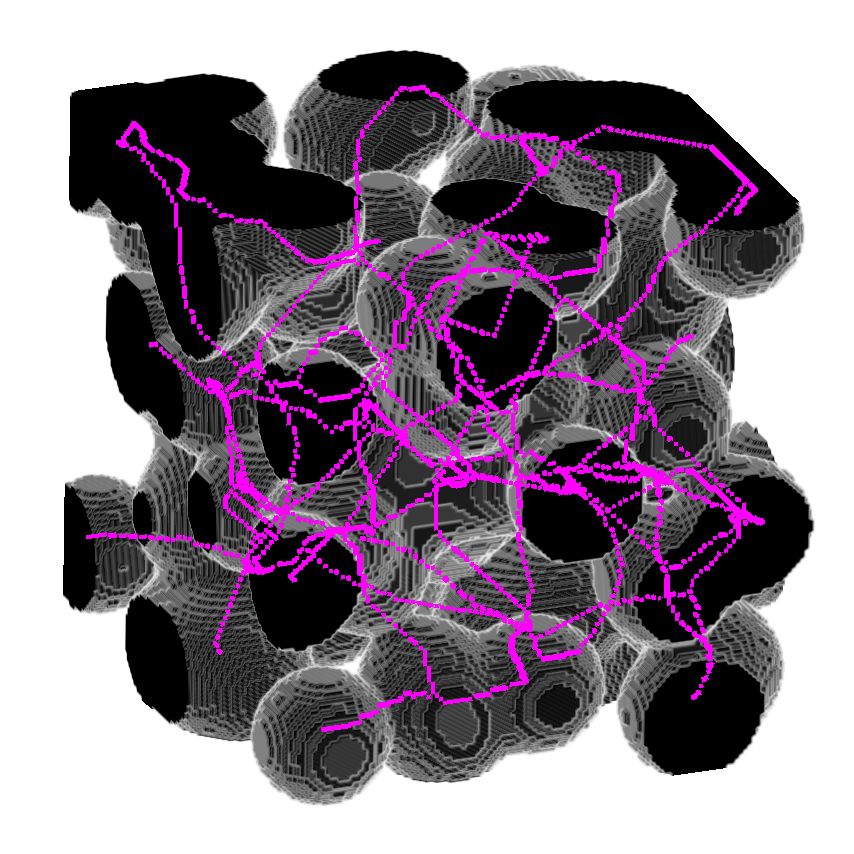
>>> viewer.add_image(image,
>>> rendering = "attenuated_mip",
>>> attenuation = 0.5,
>>> scale = [1, 1, 1])
>>> viewer.add_image(skeleton,
>>> interpolation3d = "nearest",
>>> colormap = "magenta",
>>> scale = [1, 1, 1])
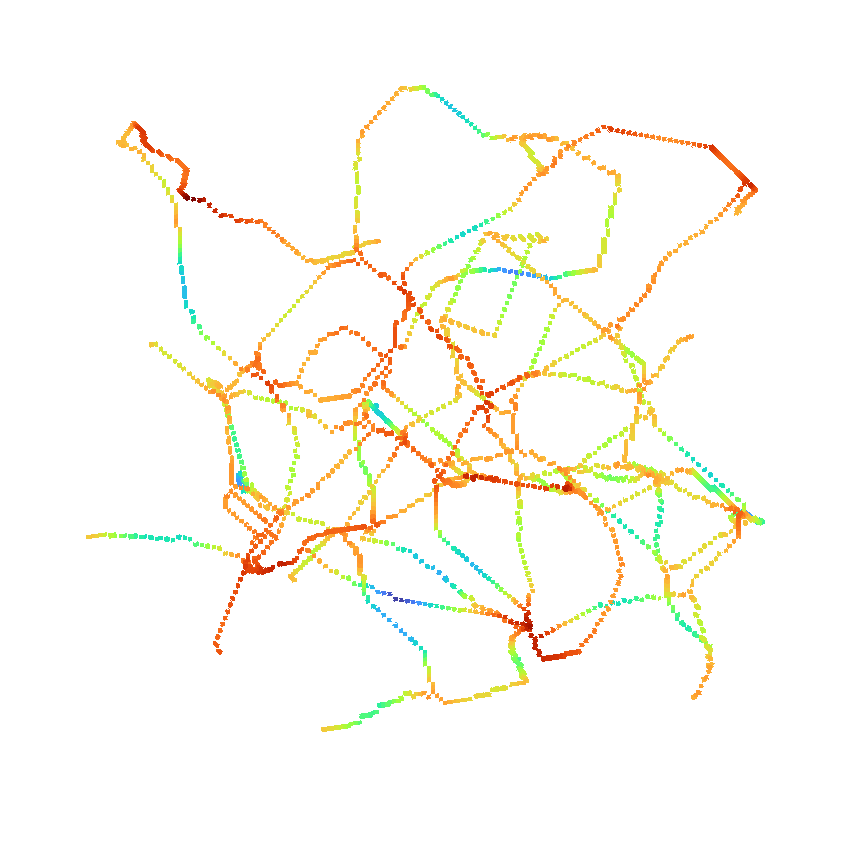
>>> napari.run()or use it to return the distance as well.
>>> import numpy as np
>>> import skimage as ski
>>> import medialaxis3d
>>> import napari
>>> rng = np.random.default_rng(1278)
>>> image = ski.data.binary_blobs(length = 128,
blob_size_fraction = 0.2,
n_dim = 3,
volume_fraction = 0.6,
rng = rng)
>>> skeleton, distance = medialaxis3d.medial_axis_3d(image,
>>> return_distance = True,
>>> size = 8,
>>> rng = rng)
>>> viewer = napari.Viewer()
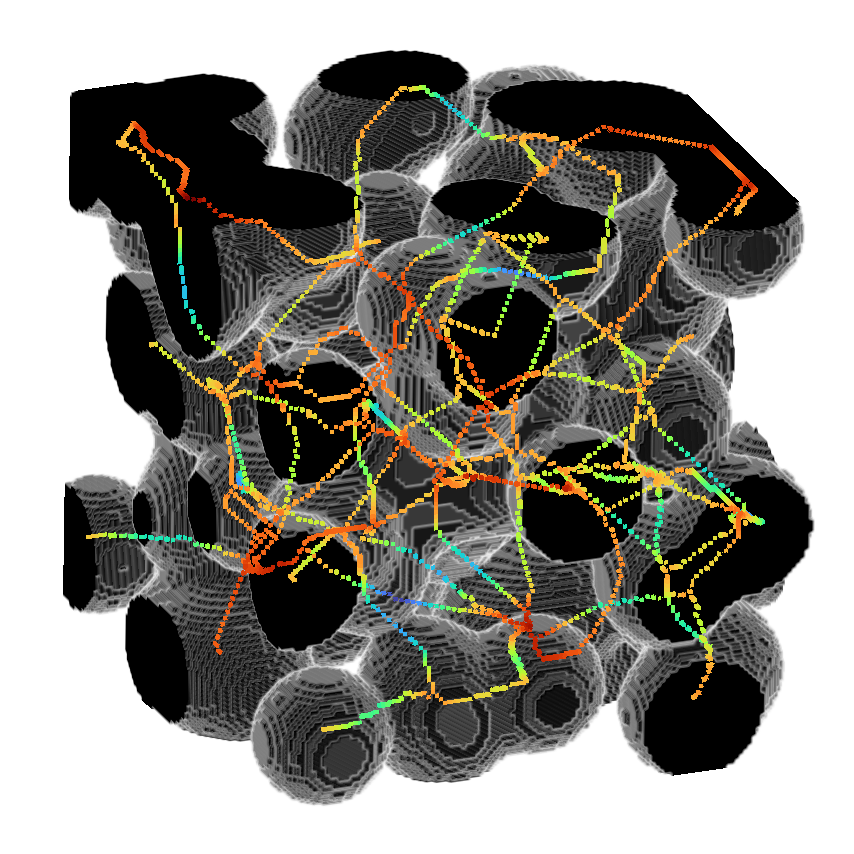
>>> viewer.add_image(image,
>>> rendering = "attenuated_mip",
>>> attenuation = 0.5,
>>> scale = [1, 1, 1])
>>> viewer.add_image(skeleton*distance,
>>> interpolation3d = "nearest",
>>> colormap = "turbo",
>>> scale = [1, 1, 1])
>>> napari.run()