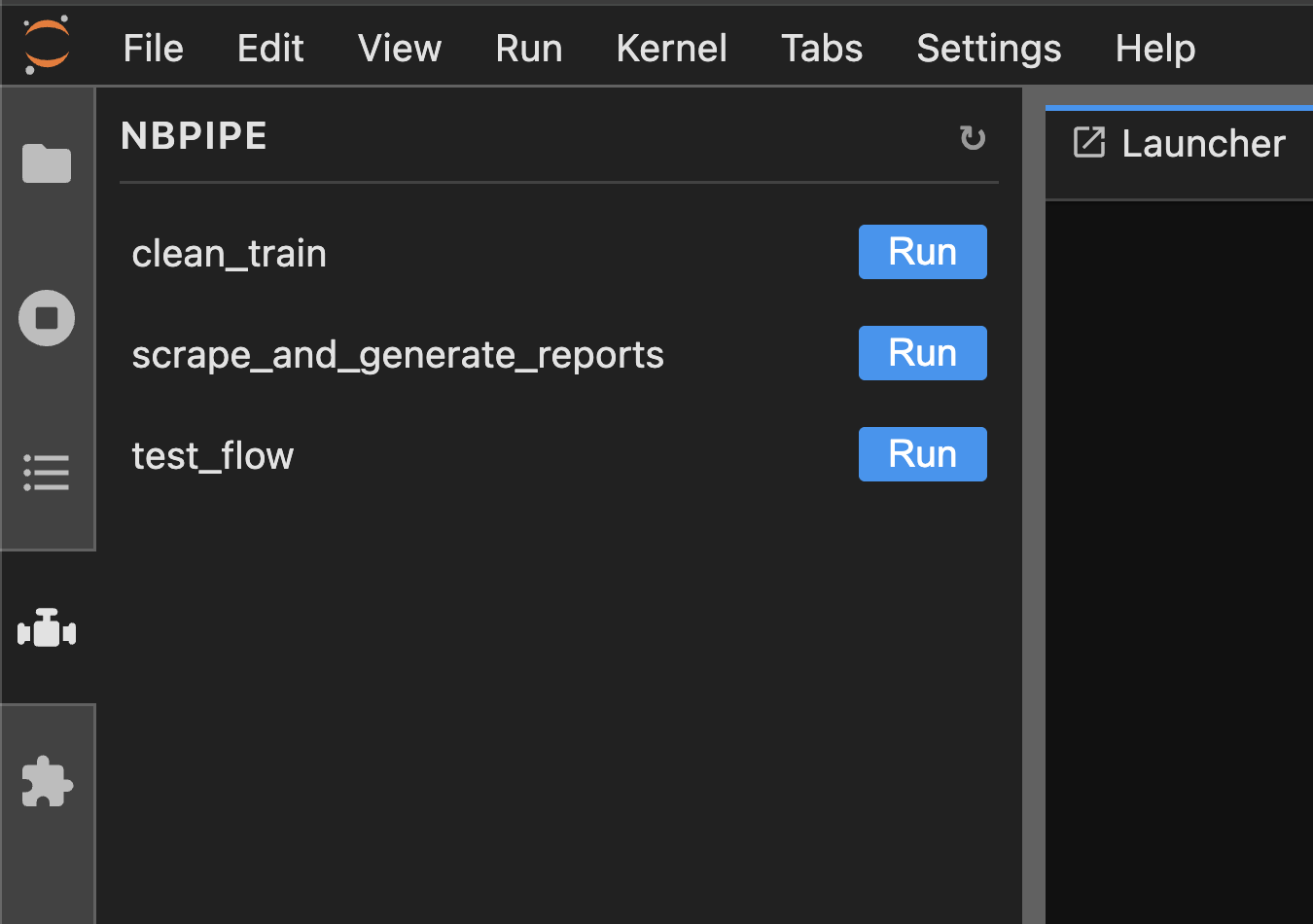
Run sequences of Jupyter notebooks as a workflow from the command line or the JupyterLab sidebar.
pip install nbpipeCreate a .nbpipe/ directory at the root of your project and add a workflow YAML file inside it.
# .nbpipe/my_workflow.yaml
# Notebook paths are relative to the project root
name: my_workflow
steps:
- notebook: prepare_data.ipynb
output: data/processed.csv
- notebook: train_model.ipynbRun it from the CLI:
nbpipe run .nbpipe/my_workflow.yamlor from JupyterLab:
| Field | Required | Description |
|---|---|---|
notebook |
yes | Notebook to run, relative to the project root |
output |
no | File the notebook must produce — workflow fails if absent |