Batch integration data#355
Conversation
…leCellOpenProblems into batch-integration-data
 LuckyMD
left a comment
LuckyMD
left a comment
There was a problem hiding this comment.
A few comments on the order of filtering in the immune data. Also, shouldn't this PR also include the task dataset functions from the batch integration graph task? In those, you should assign adata.X = adata.layers['log_scran'] again.
|
I guess the PR should also be tested... |
…leCellOpenProblems into batch-integration-data
2e569eb to
099aacf
Compare
Codecov Report
@@ Coverage Diff @@
## main #355 +/- ##
==========================================
+ Coverage 91.48% 91.86% +0.37%
==========================================
Files 83 89 +6
Lines 1973 2064 +91
Branches 117 118 +1
==========================================
+ Hits 1805 1896 +91
Misses 125 125
Partials 43 43
Flags with carried forward coverage won't be shown. Click here to find out more.
Continue to review full report at Codecov.
|
…leCellOpenProblems into batch-integration-data
|
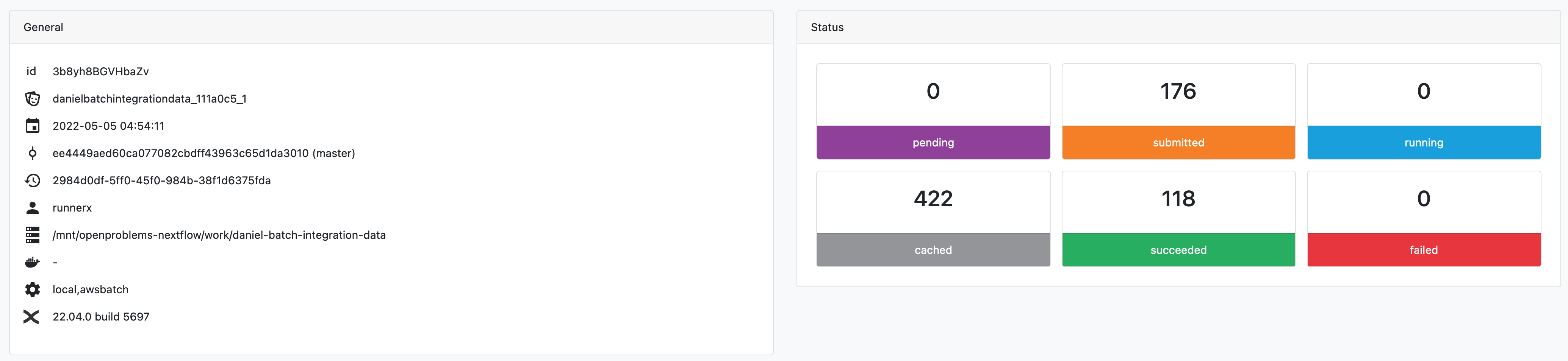
So this looks like it's all succeeded? The tower.nf logs strangely show the nextflow pipeline is still running, despite everything being complete. |
|
That's weird, |
|
I'm looking into it, I think it's a nextflow bug (that I might have just fixed.) |
|
The issue seems to persist, still running for 5 hours |
|
Looking back through the logs, it looks like this only started happening on
the self-hosted runner. Trying a reversion back to GH Actions for now.
…On Thu, May 5, 2022 at 10:38 AM Daniel Strobl ***@***.***> wrote:
[image: Screenshot 2022-05-05 at 16 32 47]
<https://user-images.githubusercontent.com/50872326/166947775-357b903d-f489-4ede-a303-88b345b47b32.png>
—
Reply to this email directly, view it on GitHub
<#355 (comment)>,
or unsubscribe
<https://github.com/notifications/unsubscribe-auth/AUHCMAUI33SNA7IYC4WGSJTVIPMNJANCNFSM5TKEP2PQ>
.
You are receiving this because you commented.Message ID:
***@***.***>
|
…leCellOpenProblems into batch-integration-data
 LuckyMD
left a comment
LuckyMD
left a comment
There was a problem hiding this comment.
few questions about the documentation and the testing that matches it, otherwise this is ready to go :). Nice job!
|
|
||
| The `openproblems-python-batch-integration` docker container is used for the methods that | ||
| can be installed without package conflicts. For R methods, the `openproblems-r-extras` | ||
| container is used. |
There was a problem hiding this comment.
Need to add other docker containers here, or make a note to add them in the other, larger PR.
| * `full_scaled` | ||
| * `hvg_scaled` | ||
|
|
||
| An example script can be found [here](methods/_example.py) |
There was a problem hiding this comment.
what is this example script meant to be?
| def check_dataset(adata): | ||
| """Check that dataset output fits expected API.""" | ||
|
|
||
| assert "X_uni" in adata.obsm |
There was a problem hiding this comment.
I assume this is old code and should now be a test for adata.layers['counts']?
There was a problem hiding this comment.
Otherwise, this is not stated in the README and should be added... it seems to be used as a representation below for the sample_method function.
|
|
||
| def check_method(adata): | ||
| """Check that method output fits expected API.""" | ||
| assert "connectivities" in adata.obsp |
There was a problem hiding this comment.
Either check for "distances" as well, or remove this from the API description in the README.
| * `adata.layers['counts']` with raw, integer UMI count data, and | ||
| * `adata.obsp['uni_connectivities']` with an unintegrated connectivity matrix generated | ||
| by `scanpy.pp.neighbors()` | ||
| * `adata.X` with log-normalized data |
There was a problem hiding this comment.
need to add adata.obsm['X_uni'] as the PCA embedding of the unintegrated representation here.
…leCellOpenProblems into batch-integration-data
|
Woohoo! |
|
Nice :). And by the deadline :D |
* Update with to accomodate multimodal data * rename bash script
* Update with to accomodate multimodal data * rename bash script Former-commit-id: 3a811a7
Add immune cell dataset from Luecken et al., 2022 for batch integration task.
Submission type
Testing
Submission guidelines
PR review checklist
This PR will be evaluated on the basis of the following checks:
__init__.pyand were tested in the pipeline