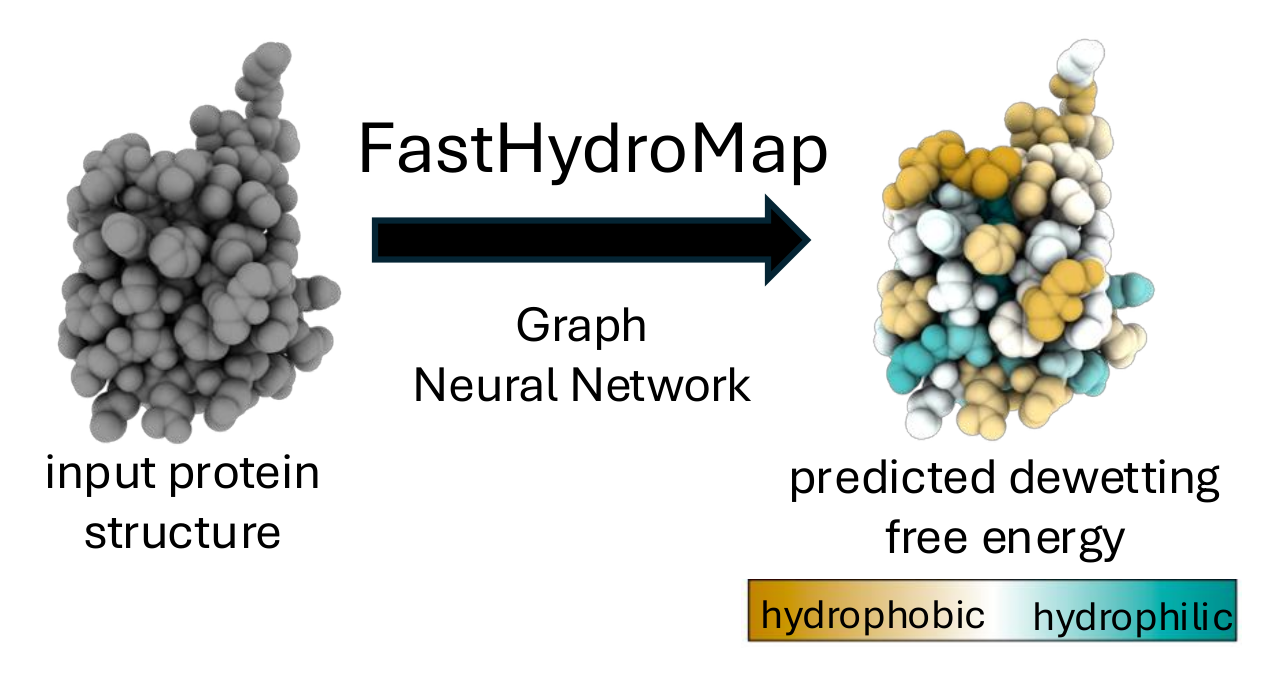
FastHydroMap predicts per-residue dewetting free energies (Fdewet) from protein structures and trajectories.
Use a fresh Python environment. Python 3.11 to 3.14 are supported.
pip install fasthydromap
fasthydromap install-torch
fasthydromap predict your_structure.pdb -o outputs/your_structure_fdewetfasthydromap install-torch defaults to the CPU build, which is usually the right choice for current FastHydroMap workloads because SASA preprocessing dominates runtime.
Advanced installation options, Docker usage, GPU Torch variants, and release workflows are documented in INSTALL.md and PYPI_RELEASE.md.
FastHydroMap supports:
- Single protein structures in
PDBformat - Protein trajectories in
DCDorXTCformat together with a matching topologyPDB
Typical usage:
# Single structure
fasthydromap predict examples/1A1U.pdb -o outputs/1A1U_fdewet
# Trajectory
fasthydromap predict-trajectory examples/proteinG.pdb examples/proteinG_short.dcd -o outputs/proteinG_fdewetFor a single structure, FastHydroMap writes:
*.csv: one row per residue withFdewet; with--parts, intrinsic and context columns are included*.pdb: a copy of the input structure with predictedFdewetwritten to B-factors
For a trajectory, FastHydroMap writes wide CSV files containing one row per frame and one column per residue.
Use --parts to also write intrinsic, context, and per-frame summary CSVs.
FastHydroMap was trained on structured single-chain proteins and the 20 canonical amino-acid chemistries. Predictions for PTMs and other non-canonical chemistries should be treated cautiously.
FastHydroMap writes Fdewet values to the B-factor column of output PDBs, so you can color structures directly in molecular viewers.
ChimeraX:
color bfactor range 4,6.5 palette ^lipophilicityPyMOL:
spectrum b, red_white_blue, minimum=4, maximum=6.5For dynamic hydrophobicity visualization in a MD trajectory, see the teaching-oriented example script
scripts/chimerax_fdewet_trajectory_example.py with a ChimeraX implementation you can adjust.
If you use FastHydroMap in your research, please cite the software release:
Lobo, S. FastHydroMap (Version 0.1.2) [Computer software]. Zenodo. https://doi.org/10.5281/zenodo.19744336
When the manuscript becomes available, please cite that as well.
Shell Lab and Shea Group