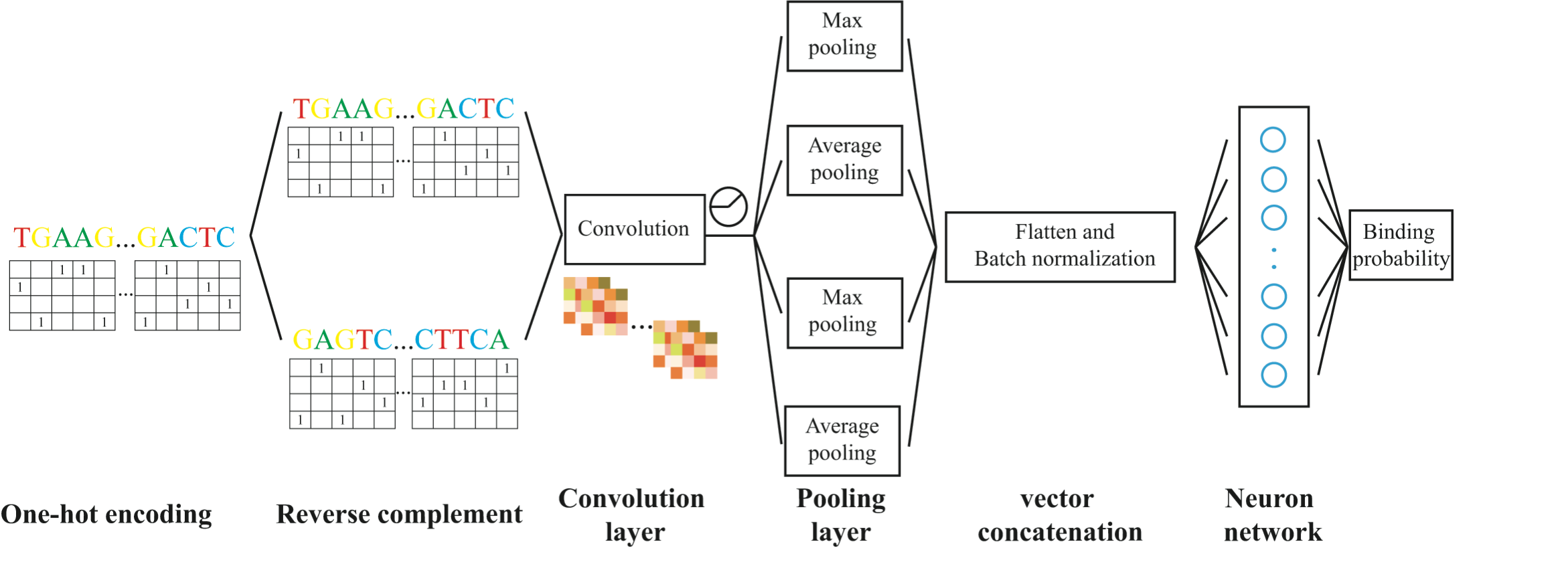
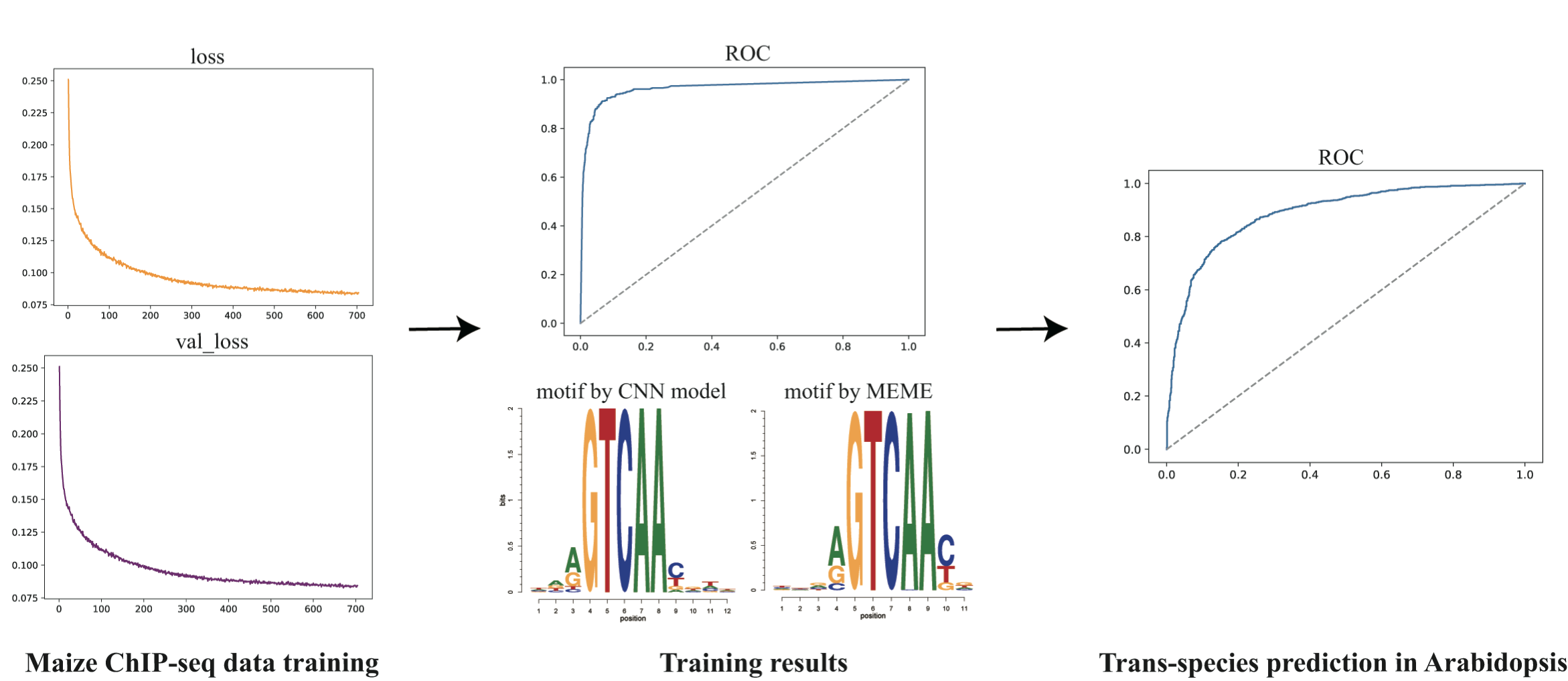
The goal of this model is to predict the binding probability of a specific transcription factor based only on DNA sequence information of the binding region by means of convolutional neural network. Below is an overview of the strcture.
tensorflow or Theano
keras
sklearn
numpy
matplotlib
tqdm
motifStack
For usage, one only needs to provide a TF narrowPeak file and genome file in FASTA format. In this demo, a narrowPeak file from maize ChIP-seq data is used.
python prepare.py zm_tf.narrowPeak zm.fa zm_tf_train.txt
python generator.py zm_tf.narrowPeak
python SeqConv.py zm_tf_train.txt
python motifGen.py zm_tf
Rscript plot_motif.r zm_tf