PETSc examples: assemble matrix analytically instead of coloring#807
Merged
PETSc examples: assemble matrix analytically instead of coloring#807
Conversation
jedbrown
commented
Sep 9, 2021
Member
|
I think my change to the solids example should be ok, but there is something strange in your PETSc branch: I think it should be a quick fix though |
Member
Author
|
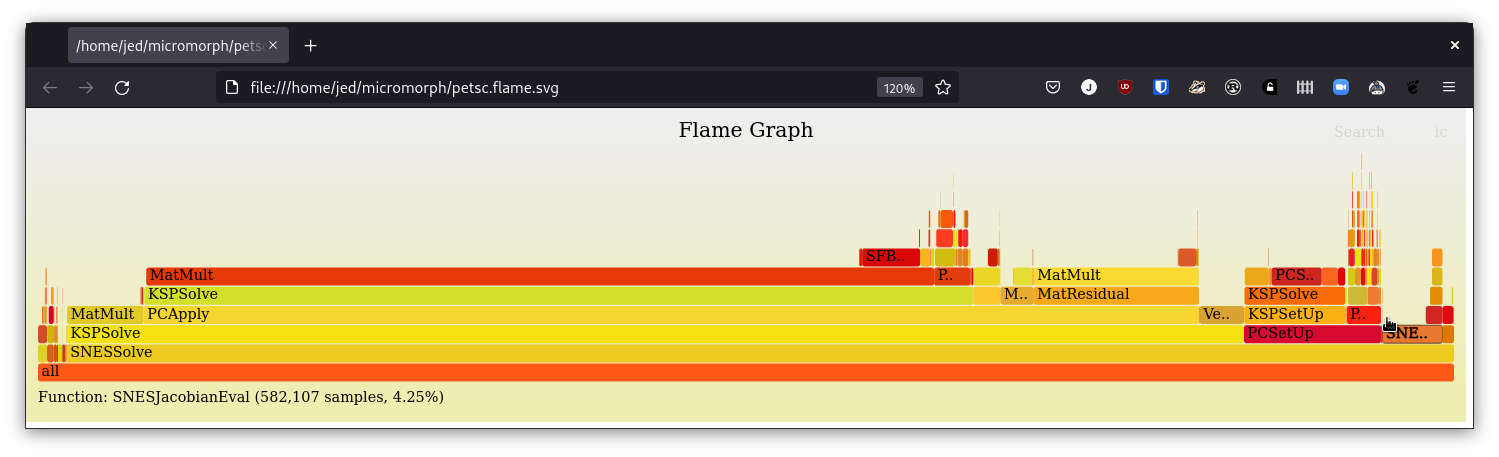
Hmm, I fixed your uninitialized variable (pushed) and it's running: |
Member
|
Still not working on my local machine. What do you get with |
Member
Author
|
Member
|
Strange. I'm on |
Member
Author
fb87e72 to
28c91eb
Compare
f1a5fd6 to
dd898e9
Compare
jeremylt
approved these changes
Jan 31, 2022
Member
 jeremylt
left a comment
jeremylt
left a comment
There was a problem hiding this comment.
Passes locally when rebased with main.
jeremylt
reviewed
Jan 31, 2022
…oring This currently requires development PETSc to correctly handle negative indices through the LocalToGlobalMapping.
d07a1cf to
cffe6a5
Compare
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
Sign up for free
to join this conversation on GitHub.
Already have an account?
Sign in to comment
Add this suggestion to a batch that can be applied as a single commit.This suggestion is invalid because no changes were made to the code.Suggestions cannot be applied while the pull request is closed.Suggestions cannot be applied while viewing a subset of changes.Only one suggestion per line can be applied in a batch.Add this suggestion to a batch that can be applied as a single commit.Applying suggestions on deleted lines is not supported.You must change the existing code in this line in order to create a valid suggestion.Outdated suggestions cannot be applied.This suggestion has been applied or marked resolved.Suggestions cannot be applied from pending reviews.Suggestions cannot be applied on multi-line comments.Suggestions cannot be applied while the pull request is queued to merge.Suggestion cannot be applied right now. Please check back later.
This currently requires development PETSc to correctly handle negative indices through the LocalToGlobalMapping.