Describe a schema in an R object#344
Conversation
|
This pull request has been linked to Shortcut Story #13273: Wrap ArraySchema as high-level object. |
9f9f353 to
a4bf68e
Compare
847a7a8 to
cf154e1
Compare
|
Some more work here to make it more like the Python sibbling that describes in array schema in 'code'. A new function This is now mostly feature complete, I have don't full round-trips yet to see if all ascii representations of enums of mapping fully (there may be some cases of, say, "ASCII" != "TILEDB_ASCII"). For example for the Penguins array (with NAs) we now get this: > library(tiledb)
TileDB R 0.10.2 with TileDB Embedded 2.7.0. See https://tiledb.com for more information.
> arr <- tiledb_array("/tmp/tiledb/penguins/")
> describe(arr)
dims <- c(tiledb_dim(name="species", domain=c(NULL,NULL), tile=NULL, type="ASCII"),
tiledb_dim(name="island", domain=c(NULL,NULL), tile=NULL, type="ASCII")))
dom <- tiledb_domain(dims=dims)
attrs <- c(tiledb_attr(name="bill_length_mm", type="FLOAT64", ncells=1, nullable=TRUE, filter_list=c(tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))))),
tiledb_attr(name="bill_depth_mm", type="FLOAT64", ncells=1, nullable=TRUE, filter_list=c(tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))))),
tiledb_attr(name="flipper_length_mm", type="INT32", ncells=1, nullable=TRUE, filter_list=c(tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))))),
tiledb_attr(name="body_mass_g", type="INT32", ncells=1, nullable=TRUE, filter_list=c(tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))))),
tiledb_attr(name="sex", type="ASCII", ncells=NA, nullable=TRUE, filter_list=c(tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))))),
tiledb_attr(name="year", type="INT32", ncells=1, nullable=FALSE, filter_list=c(tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))))))
sch <- tiledb_array_schema(domain=dom, attrs=attrs, cell_order="COL_MAJOR", tile_order="COL_MAJOR", sparse=TRUE, capacity=10000, allow_dupes=TRUE,
coord_filters=filter_list=c(tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))))),
offset_filters=filter_list=c(tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))))))
> after which |
 johnkerl
left a comment
johnkerl
left a comment
There was a problem hiding this comment.
awesome @eddelbuettel !!
one little fine-tune opportunity, i ran this for a few arrays then vim'ed the result & i am seeing some parenthesis imbalances --
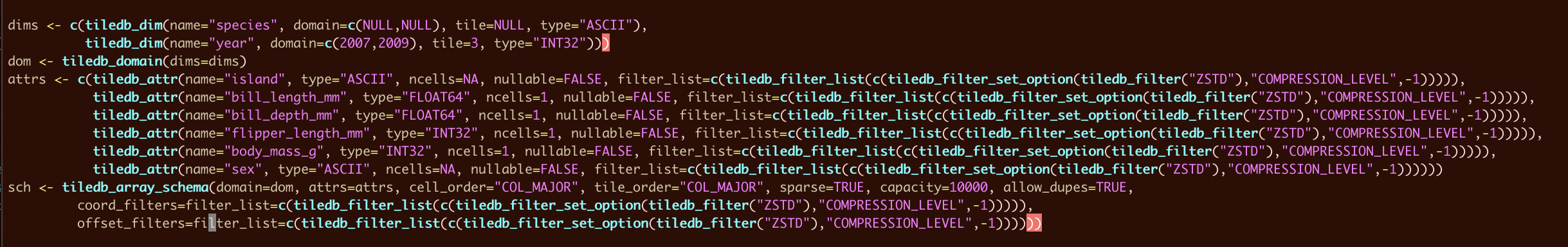
to be discussed if this should be an option to show() instead
367b0ad to
1a9379c
Compare
|
We all may need to gab a little in a little while (as per my chat with @ihnorton) as we a) probably want to restore the object dump from core as an optional feature and b) need to work out if we want these verbose code pretty printers or the "look like core but ain't" you added as default as two may seem like one too many. Not urgent, but one of those things where a little chat prior to unannounced PRs can work wonders. Also note that the code in the PR so far 'only looks pretty' but hasn't been to the dance yet. I haven't done any round-robin tests yet. I am sure there may be a many-legged creature be hiding in a corner or two. |
This PR gather schema information we can use to both print schema creation command, and to summarize array objects directly for more fine-grained formatting. It is a rather unfortunate this only comes together now as @johnkerl could most likely have saved some time had I put this together earlier.
I will leave this as a draft for now. It currently returns a list with two data frames for array (high-level) descriptives and then one for all 'data' columns. As dimensions and attributes are in fact a little distinct it may be beneficial to return one each for dimensions and attributes.
Current output format showing the two data frames directly on two sample arrays: