Conversation
|
This pull request has been linked to Shortcut Story #13744: Update show methods to more compact 'schema creation style' used by tiledb-py. |
|
Awesome! Let me give this a spin in Jupyter just as a road-test. :) |
|
OK, test-drove in Jupyter! My thinking (perhaps overwrought) was to make sure if someone prints the entire schema, it comes out OK -- and likewise if they just print the domain, etc. Full code: Some notes:
Also I would add that I love that the output from |
 johnkerl
left a comment
johnkerl
left a comment
There was a problem hiding this comment.
Review comments here (sorry if it's confusing to narrate through a screenshot)
#355 (comment)
I am not sure what you are trying to say here. We do have show methods for
Ahh. Likely old TileDB Core instance so no validity filter. Will try to fix.
Sorry can you be more specific? Where? Can you maybe show it more directly than "somewhere in those X lines in a screenshot" ? Also I ran the code generated from a schema and that worked. (Of course, no running example proofs anything about other possible bugs...)
These calls seem "wrong". Maybe I need to add type checkers: > flt1 <- tiledb_filter("DOUBLE_DELTA")
> flt2 <- tiledb_filter("CHECKSUM_SHA256")
> flt3 <- tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",4)
> show(flt1)
tiledb_filter("DOUBLE_DELTA")
> show(flt2)
tiledb_filter("CHECKSUM_SHA256")
> show(flt3)
tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",4)
> fltlst <- tiledb_filter_list(c(flt1, flt2, flt3))
> show(fltlst)
tiledb_filter_list(c(tiledb_filter("DOUBLE_DELTA")), tiledb_filter("CHECKSUM_SHA256")), tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",4)))
> dims <- tiledb_dim(name="rows", domain=c(1L,40L), tile=40L, type="INT32")
> show(dims)
tiledb_dim(name="rows", domain=c(1L,40L), tile=40L, type="INT32")
> (As an aside, the debugging in the notebook may be more cumbersome. The code works the same within/without.) The PR also makes no claim to be adding |
|
@johnkerl The trailing comma in case of less than three filter lists is taken care of (and tested against 2.5.3 too). I found no extra paren so no change there. |
|
Sorry for the confusion @eddelbuettel !
My apologies; it looks like the Re imbalanced parens, here's a screenshot of the top of the third gist in vim -- appears to be simply an extra final |
|
Ok, that I can reproduce. Minimal example: > uri <- "tiledb://TileDB-Inc/gtex-analysis-rnaseqc-gene-tpm"
> arr <- tiledb_array(uri)
> sch <- schema(arr)
> sch
tiledb_array_schema(
domain=tiledb_domain(c(tiledb_dim(name="gene_id", domain=c(NULL,NULL), tile=NULL, type="ASCII"), tiledb_dim(name="sample", domain=c(NULL,NULL), tile=NULL, type="ASCII"))),
attrs=c(tiledb_attr(name="tpm", type="FLOAT64", ncells=1, nullable=FALSE, filter_list=tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",1))))),
cell_order="ROW_MAJOR", tile_order="ROW_MAJOR", capacity=18000, sparse=TRUE, allows_dups=FALSE,
coords_filter_list=tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))),
offsets_filter_list=tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("POSITIVE_DELTA"),"POSITIVE_DELTA_MAX_WINDOW",1024)), tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",1))),
validity_filter_list=tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("RLE"),"COMPRESSION_LEVEL",-1)))
)
>
>
> fl <- filter_list(sch)
> fl
$coords
tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1)))
$offsets
tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("POSITIVE_DELTA"),"POSITIVE_DELTA_MAX_WINDOW",1024)), tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",1)))
$validity
tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("RLE"),"COMPRESSION_LEVEL",-1)))
> I'll work that out tomorrow to correct the |
|
Looks like it is sorted out: |
|
If you have some TileDB arrays on disk you can use this ad-hoc script to extract the executable schema, and re-create it. #!/bin/bash
if [ $# -lt 1 ]; then
echo "Usage: $0 uri"
exit 1
fi
tf=$(mktemp)
echo "library(tiledb)" > ${tf}
echo -n "tiledb_array_create(uri=tempfile()," >> ${tf}
# use littler: r -ltiledb -e"show(schema(tiledb_array(\"$1\")))" >> ${tf}
# or Rscript: Rscript -e 'suppressMessages(library(tiledb)); show(schema(tiledb_array(\"$1\")))" >> ${tf}
Rscript -e "suppressMessages(library(tiledb)); show(schema(tiledb_array(\"$1\")))" >> ${tf}
echo ")" >> ${tf}
echo "cat(\"Done!\\n\")" >> ${tf}
cat ${tf}
Rscript ${tf}
rm -v ${tf} |
|
@johnkerl The > library(tiledb)
TileDB R 0.10.2 with TileDB Embedded 2.7.0. See https://tiledb.com for more information.
> uri <- "tiledb://TileDB-Inc/gtex-analysis-rnaseqc-gene-tpm"
> arr <- tiledb_array(uri, query_type="READ", as.data.frame=TRUE)
> sch <- schema(arr)
> cat("SCHEMA\n")
SCHEMA
> show(sch)
tiledb_array_schema(
domain=tiledb_domain(c(tiledb_dim(name="gene_id", domain=c(NULL,NULL), tile=NULL, type="ASCII"), tiledb_dim(name="sample", domain=c(NULL,NULL), tile=NULL, type="ASCII"))),
attrs=c(tiledb_attr(name="tpm", type="FLOAT64", ncells=1, nullable=FALSE, filter_list=tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",1))))),
cell_order="ROW_MAJOR", tile_order="ROW_MAJOR", capacity=18000, sparse=TRUE, allows_dups=FALSE,
coords_filter_list=tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1))),
offsets_filter_list=tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("POSITIVE_DELTA"),"POSITIVE_DELTA_MAX_WINDOW",1024), tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",1))),
validity_filter_list=tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("RLE"),"COMPRESSION_LEVEL",-1)))
)
> fl <- filter_list(sch)
> cat("\nFILTER_LIST\n")
FILTER_LIST
> show(fl)
$coords
tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",-1)))
$offsets
tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("POSITIVE_DELTA"),"POSITIVE_DELTA_MAX_WINDOW",1024), tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",1)))
$validity
tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("RLE"),"COMPRESSION_LEVEL",-1)))
> dom <- domain(sch)
> cat("\nDOMAIN\n")
DOMAIN
> show(dom)
tiledb_domain(c(tiledb_dim(name="gene_id", domain=c(NULL,NULL), tile=NULL, type="ASCII"), tiledb_dim(name="sample", domain=c(NULL,NULL), tile=NULL, type="ASCII")))
> ndim <- tiledb_ndim(dom)
> cat("\nNDIM\n")
NDIM
> show(ndim)
[1] 2
> dims <- dimensions(dom)
> cat("\nDIM1\n")
DIM1
> show(dims[[1]])
tiledb_dim(name="gene_id", domain=c(NULL,NULL), tile=NULL, type="ASCII")
> cat("\nDIM2\n")
DIM2
> show(dims[[2]])
tiledb_dim(name="sample", domain=c(NULL,NULL), tile=NULL, type="ASCII")
> cat("\nDIM1 FL\n")
DIM1 FL
> show(filter_list(dims[[1]]))
> cat("\nDIM2 FL\n")
DIM2 FL
> show(filter_list(dims[[2]]))
> nattr <- length(attrs(sch))
> cat("\nNATTR\n")
NATTR
> show(nattr)
[1] 1
> attr1 <- attrs(sch, 1)
> cat("\nATTR1\n")
ATTR1
> show(attr1)
tiledb_attr(name="tpm", type="FLOAT64", ncells=1, nullable=FALSE, filter_list=tiledb_filter_list(c(tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",1))))
> fla <- filter_list(attr1)
> cat("\nATTR1 FL\n")
ATTR1 FL
> show(fla[0])
tiledb_filter_set_option(tiledb_filter("ZSTD"),"COMPRESSION_LEVEL",1)
> |
 johnkerl
left a comment
johnkerl
left a comment
There was a problem hiding this comment.
Awesome @eddelbuettel !! :D
#355 (comment) is really the bomb -- built-in syntax checking + user empowerment FTW! :D
|
"It takes a village" 😂 -- started drafting the NEWS for 0.11.0 and we about 'large number' of PRs on this topic but I like the place we got, even if it some iterations! |
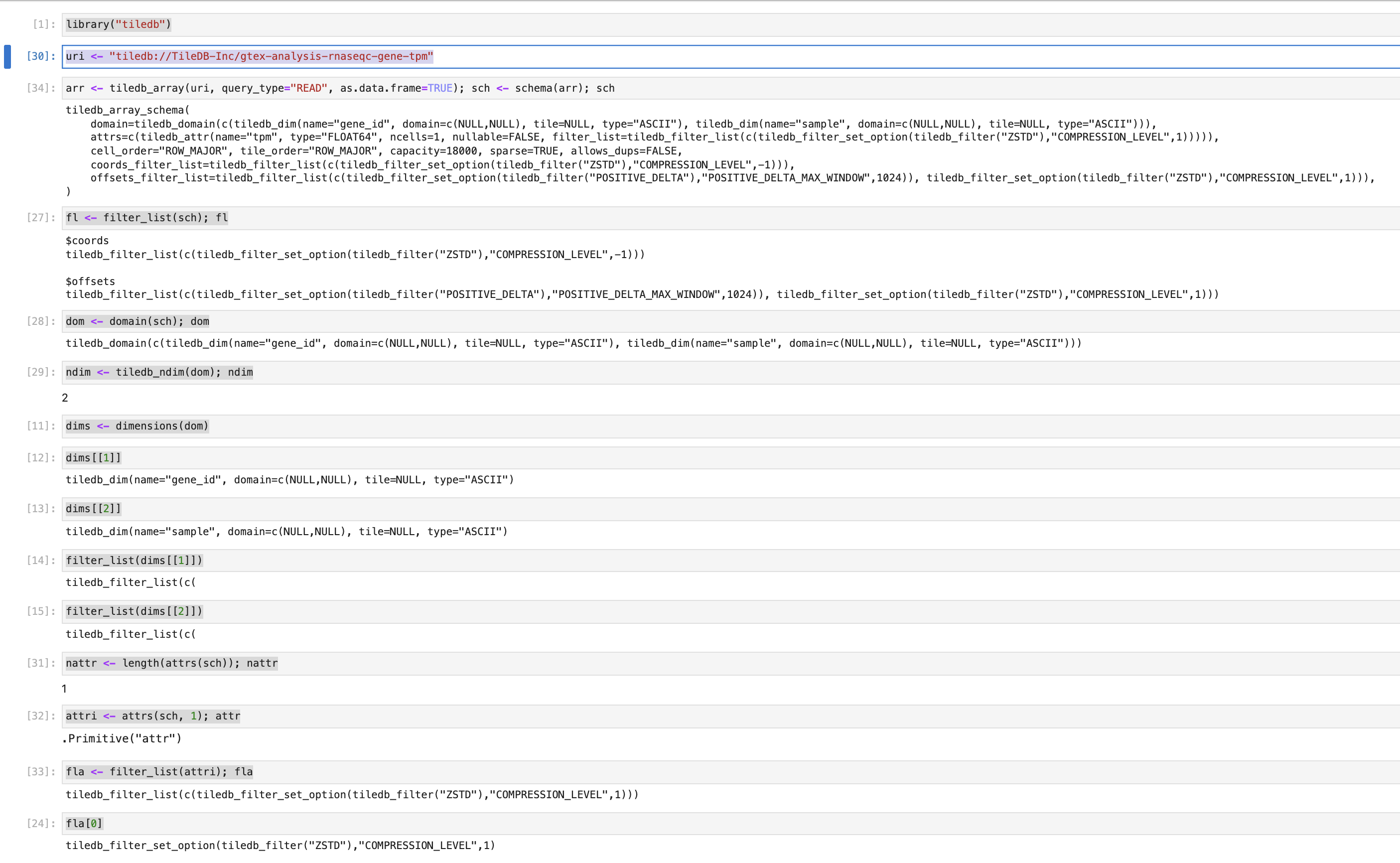

This PR updates the newer
show()methods. They now work better in isolation as well as when composed, are more compact and a little closer to the Python equivalents.