Conversation
|
Hello Maria. This PR is not working on RTD, the compilation is failing. |
|
it might be useful to enable PR builds on readthedocs: https://docs.readthedocs.io/en/stable/pull-requests.html That way you get feedback on each PR if it can be successfully built and also get a preview of the new documentation if it was successful. |
|
Thanks @arthurPignetOwkin , indeed RTD didn't see the data path. I added a selection that if the path is not found the data file is loaded directly from github. It should work now |
|
Hi, thanks a lot for your work @maikia. For some reason my RTD account can't be added to the pydeseq2 project. I've contacted the RTD support on this issue, there should be a fixed released this week. Once this is fixed I should be able to review your PR. |
|
"PYDESEQ2 PIPELINE" title is too big it's threatening. |
ops, sorry. updated comment: I though you were referring to the title of the documentation and not the notebook. I will include this review comment in the revision. Thanks for the suggestion |
b11408d to
044a5a2
Compare
|
Hi @maikia, I rebased |
 BorisMuzellec
left a comment
BorisMuzellec
left a comment
There was a problem hiding this comment.
Thanks for the nice work @maikia, the example looks great!
I've just suggested a few changes to the contents of the example itself: I don't think it's relevant to mention TCGA data (since we're not giving precise instructions to download it) nor to keep the SAVE flag and output pickling. Let me know what you think about this.
| dds = DeseqDataSet( | ||
| counts_df, | ||
| clinical_df, | ||
| design_factors="condition" if DATASET == "synthetic" else "high_grade", |
There was a problem hiding this comment.
| design_factors="condition" if DATASET == "synthetic" else "high_grade", | |
| design_factors="condition", |
There was a problem hiding this comment.
again, I think this should be explained, not removed?
There was a problem hiding this comment.
Not sure, the high_grade field is specific to TCGA data. We could add a comment saying that "condition" is a column of clinical_df
|
@BorisMuzellec Perhaps we should re-discuss the purpose of this example? In view of your comments it seems that we had a different idea :-) |
 BorisMuzellec
left a comment
BorisMuzellec
left a comment
There was a problem hiding this comment.
@maikia I've updated my review following our conversation: let's keep the gene-filtering and output-saving parts, but remove TCGA.
|
Thanks @BorisMuzellec for your review. |
…tly from github (rebase and fix conflict with load_example_data refactoring)
Co-authored-by: Boris Muzellec <BorisMuzellec@users.noreply.github.com>
Co-authored-by: Boris Muzellec <BorisMuzellec@users.noreply.github.com>
Co-authored-by: Boris Muzellec <BorisMuzellec@users.noreply.github.com>
Co-authored-by: Boris Muzellec <BorisMuzellec@users.noreply.github.com>
 BorisMuzellec
left a comment
BorisMuzellec
left a comment
There was a problem hiding this comment.
Hi @maikia, thanks a lot for the great work!
I agree that there is still some work to do to improve the example (we should add further explanations) but I think we should merge this PR, which already implements the notebook rendering in the docs, and save further improvements on the text for a later PR.
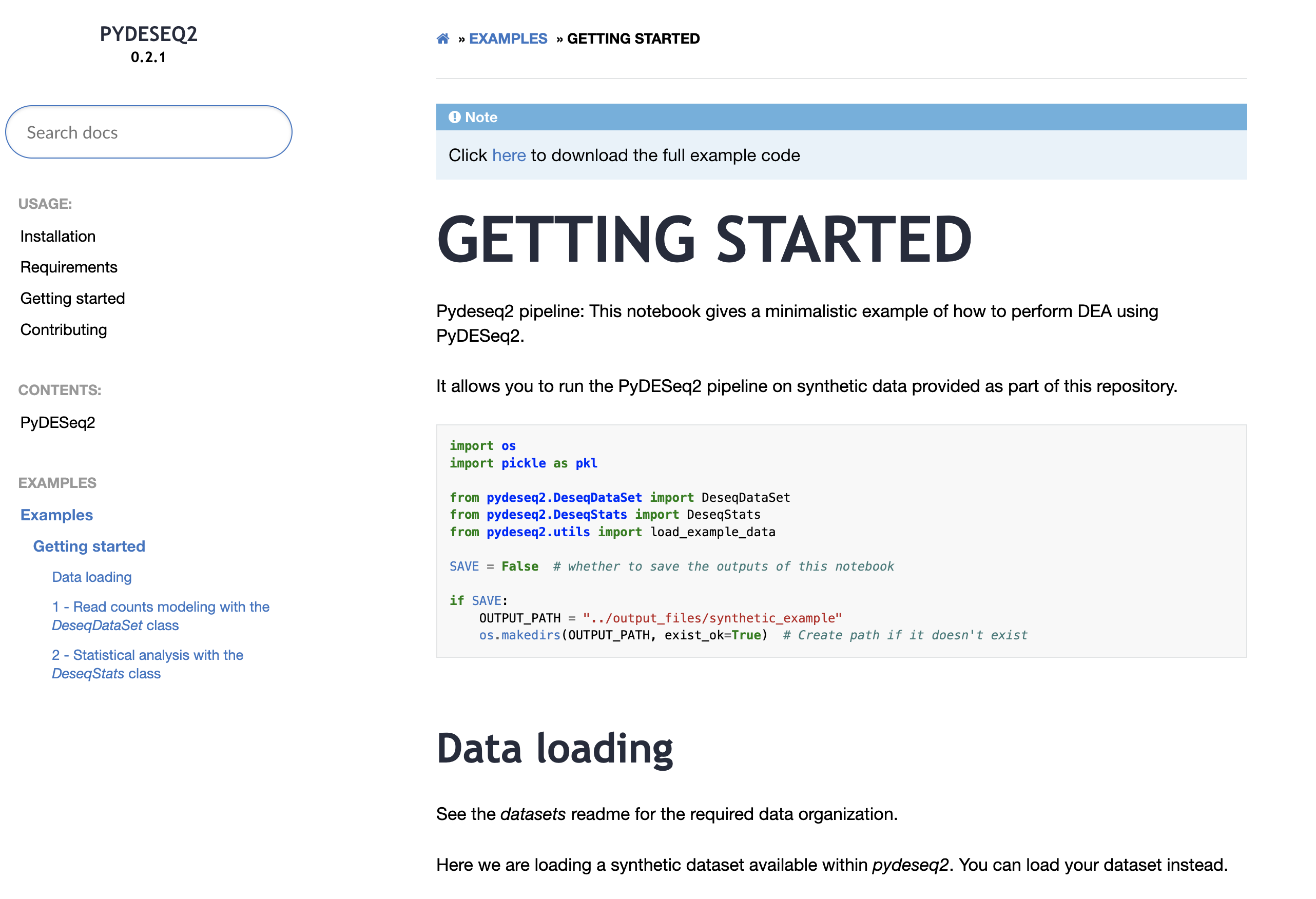
introducing sphinx-gallery with the first example converted from the notebooks folder.
Note:
Here how it looks like now (locally):